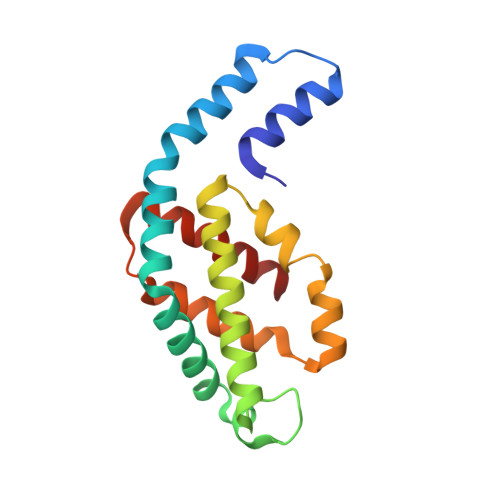
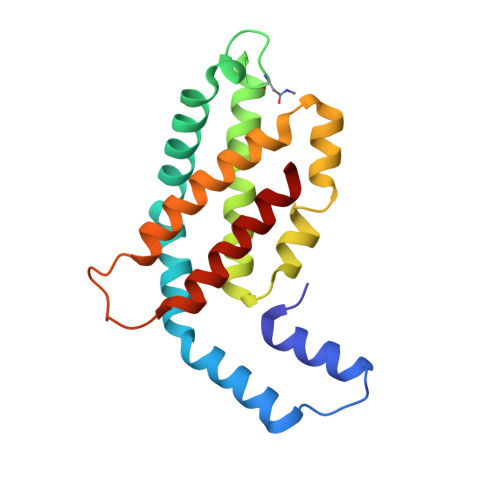
Isolation, crystallization, crystal structure analysis and refinement of constitutive C-phycocyanin from the chromatically adapting cyanobacterium Fremyella diplosiphon at 1.66 A resolution.
Duerring, M., Schmidt, G.B., Huber, R.(1991) J Mol Biology 217: 577-592
- PubMed: 1899708 Search on PubMed
- DOI: https://doi.org/10.1016/0022-2836(91)90759-y
- Primary Citation Related Structures:
1CPC - PubMed Abstract:
Constitutive phycocyanin from cyanobacterium Fremyella diplosiphon (Calothrix sp. PCC 7601) grown in green light, has been isolated and crystallized. The crystals belong to the space group R3 with cell constants a = b = 180.26 A, c = 61.24 A, alpha = beta = 90 degrees, gamma = 120 degrees. The crystal structure has been determined by Patterson search techniques using the molecular model of C-phycocyanin from the cyanobacterium Agmenellum quadruplicatum. The asymmetric unit of the crystal cell consists of two (alpha beta)-monomers related by a local dyad. Three asymmetric units are arranged around a crystallographic triad and form an (alpha beta)6-hexamer, the functional unit in the native antenna rod. The initial structure has been refined in a cyclic manner by energy-restrained crystallographic refinement and modelling until the conventional crystallographic R-factor converged at 18.1% with data to a resolution of 1.66 A. The molecular structure resembles closely the C-phycocyanins of Mastigocladus laminosus and A. quadruplicatum. The conformation and configuration of the alpha-84 and beta-84 chromophores is very similar to the corresponding chromophores in the trimeric C-phycocyanin of M. laminosus, whereas the beta-155 chromophore differs in configuration with C(4)-Z, C(10)-Z and C(15)-Z compared to C(4)-Z, C(10)-Z, C(15)-Z,E. The stereochemistry of the beta-155 chiral centres is C(2)-RC(3)-R and C(31)-S, respectively, whereas alpha-84 and beta-84 have C(2)-RC(3)-R and C(31)-R. The amino acid sequences of constitutive and inducible phycocyanin differ mainly in residues located on the surface of the beta-subunits that mediate the inter-hexameric contacts.
- Max-Planck-Institut für Biochemie, München, Germany.
Organizational Affiliation: