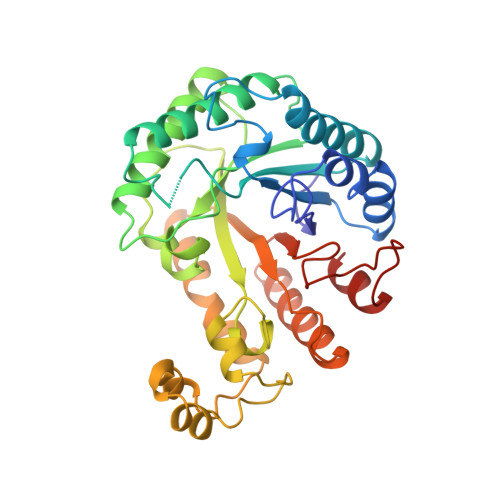
A common protein fold and similar active site in two distinct families of beta-glycanases.
Dominguez, R., Souchon, H., Spinelli, S., Dauter, Z., Wilson, K.S., Chauvaux, S., Beguin, P., Alzari, P.M.(1995) Nat Struct Biol 2: 569-576
- PubMed: 7664125 Search on PubMed
- DOI: https://doi.org/10.1038/nsb0795-569
- Primary Citation Related Structures:
1CEC, 1XYZ - PubMed Abstract:
The structure of Clostridium thermocellum endoglucanase CelC, a member of the largest cellulase family (family A), has been determined at 2.15 A resolution. The protein folds into an (alpha/beta)8 barrel, with a deep active-site cleft generated by the insertion of a helical subdomain. The structure of the catalytic core of xylanase XynZ, which belongs to xylanase family F, has been determined at 1.4 A resolution. In spite of significant differences in substrate specificity and structure (including the absence of the helical subdomain), the general polypeptide folding pattern, architecture of the active site and catalytic mechanism of XynZ and CelC are similar, suggesting a common evolutionary origin.
- Unité d'Immunologie Structurale, URA 1961 CNRS, Institute Pasteur, Paris, France.
Organizational Affiliation: