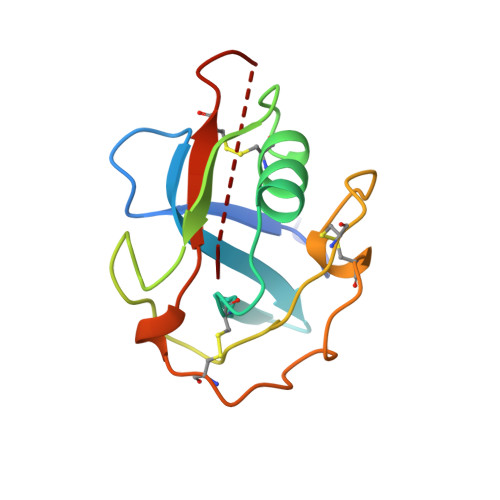
Crystal structure of a scavenger receptor cysteine-rich domain sheds light on an ancient superfamily.
Hohenester, E., Sasaki, T., Timpl, R.(1999) Nat Struct Biol 6: 228-232
- PubMed: 10074941 Search on PubMed
- DOI: https://doi.org/10.1038/6669
- Primary Citation Related Structures:
1BY2 - PubMed Abstract:
Scavenger receptor cysteine-rich (SRCR) domains are found widely in cell surface molecules and in some secreted proteins, where they are thought to mediate ligand binding. We have determined the crystal structure at 2.0 A resolution of the SRCR domain of Mac-2 binding protein (M2BP), a tumor-associated antigen and matrix protein. The structure reveals a curved six-stranded beta-sheet cradling an alpha-helix. Structure-based sequence alignment demonstrates that the M2BP SRCR domain is a valid template for the entire SRCR protein superfamily. This allows an interpretation of previous mutagenesis data on ligand binding to the lymphocyte receptor CD6.
- Department of Crystallography, Birkbeck College, University of London, UK.
Organizational Affiliation: