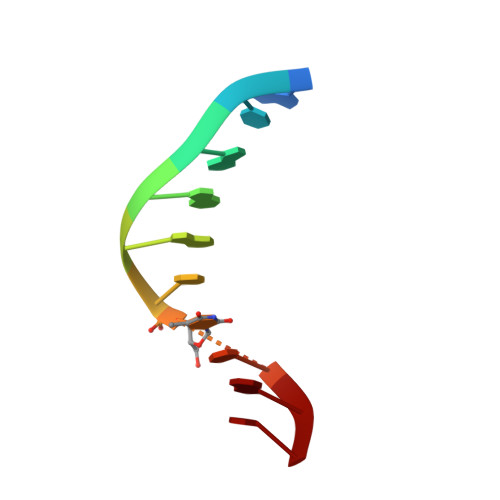
NMR solution structures of [d(GCGAAT-3'-3'-alphaT-5'-5'-CGC)2] and its unmodified control.
Aramini, J.M., Mujeeb, A., Germann, M.W.(1998) Nucleic Acids Res 26: 5644-5654
- PubMed: 9837995 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/26.24.5644
- Primary Citation Related Structures:
1BWT, 1BX5 - PubMed Abstract:
We present the high-resolution solution structures of a self-complementary DNA decamer duplex featuring a single alpha-anomeric nucleotide per strand encompassed by a set of 3'-3' and 5'-5' phosphodiester linkages, d(GCGAAT-3'-3'-alphaT-5'-5'-CGC)2, alphaT, and its unmodified control, d(GCGAATTCGC)2, obtained by restrained molecular dynamics. Interproton distance and deoxyribose ring torsion angle restraints were deduced from homonuclear NOESY and DQF-COSY data, respectively. For both the control and alphaT duplexes, excellent global convergence was observed from two different (A- and B-) starting models. The final average structures of the two duplexes are highly homologous, and overall possess the traits characteristic of right-handed B-DNA duplexes. However, localized differences between the two structures stem from the enhanced conformational exchange in the deoxyribose ring of the cytidine following the 5'-5' linkage, the C3'- exo pseudorotation phase angle of the alpha-nucleotide, and unusual backbone torsions in the 3'-3' and 5'-5' phosphodiester linkages. The structural data reported here are relevant to the design of antisense therapeutics comprised of these modifications.
- Department of Microbiology and Immunology, Kimmel Cancer Institute, Thomas Jefferson University, Philadelphia, PA 19107, USA.
Organizational Affiliation: