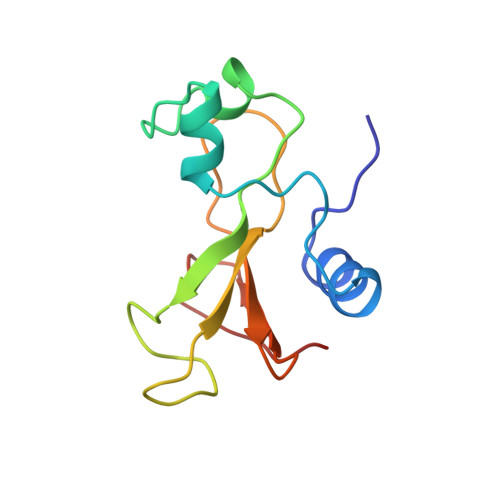
Determination of the three-dimensional solution structure of barnase using nuclear magnetic resonance spectroscopy.
Bycroft, M., Ludvigsen, S., Fersht, A.R., Poulsen, F.M.(1991) Biochemistry 30: 8697-8701
- PubMed: 1888730 Search on PubMed
- DOI: https://doi.org/10.1021/bi00099a030
- Primary Citation Related Structures:
1BNR - PubMed Abstract:
The solution conformation of the ribonuclease barnase has been determined by using 1H nuclear magnetic resonance (NMR) spectroscopy. The 20 structures were calculated by using 853 interproton distance restraints obtained from analyses of two-dimensional nuclear Overhauser spectra, 72 phi and 53 chi 1 torsion angle restraints, and 17 hydrogen-bond distance restraints. The calculated structures contain two alpha-helices (residues 6-18 and 26-34) and a five-stranded antiparallel beta-sheet (residues 50-55, 70-75, 85-91, 94-101, and 105-108). The core of the protein is formed by the packing of one of the alpha-helices (residues 6-18) onto the beta-sheet. The average RMS deviation between the calculated structures and the mean structure is 1.11 A for the backbone atoms and 1.75 A for all atoms. The protein is least well-defined in the N-terminal region and in three large loops. When these regions are excluded, the average RMS deviation between the calculated structures and the mean structure for residues 5-34, 50-56, 71-76, 85-109 is 0.62 A for the backbone atoms and 1.0 A for all atoms. The NMR-derived structure has been compared with the crystal structure of barnase [Mauguen et al. (1982) Nature (London) 297, 162-164].
- Cambridge Centre for Protein Engineering, University Chemical Laboratory, U.K.
Organizational Affiliation: