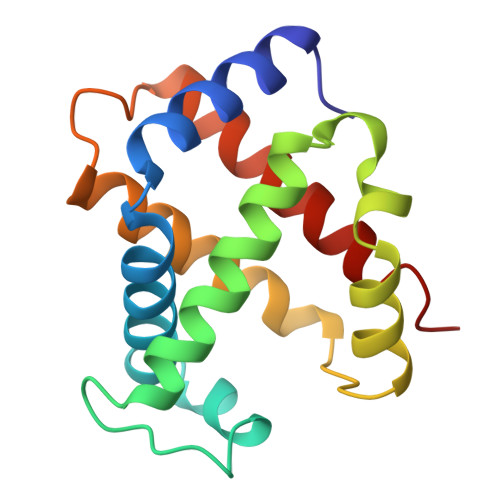
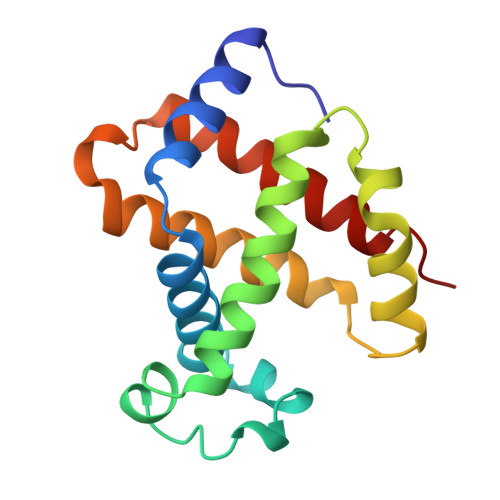
High-resolution crystal structures of human hemoglobin with mutations at tryptophan 37beta: structural basis for a high-affinity T-state,.
Kavanaugh, J.S., Weydert, J.A., Rogers, P.H., Arnone, A.(1998) Biochemistry 37: 4358-4373
- PubMed: 9521756 Search on PubMed
- DOI: https://doi.org/10.1021/bi9708702
- Primary Citation Related Structures:
1A00, 1A01, 1A0U, 1A0Z - PubMed Abstract:
The high-resolution X-ray structures of the deoxy forms of four recombinant hemoglobins in which Trp37(C3)beta is replaced with Tyr (betaW37Y), Ala (betaW37A), Glu (betaW37E), or Gly (betaW37G) have been refined and analyzed with superposition methods that partition mutation-induced perturbations into quaternary structure changes and tertiary structure changes. In addition, a new cross-validation statistic that is sensitive to local changes in structure (a "local Rfree" parameter) was used as an objective measure of the significance of the tertiary structure changes. No significant mutation-induced changes in tertiary structure are detected at the mutation site itself for any of the four mutants studied. Instead, disruption of the intersubunit contacts associated with Trp37(C3)beta results in (1) a change in quaternary structure at the alpha1beta2 interface, (2) alpha subunit tertiary structure changes that are centered at Asp94(G1)alpha-Pro95(G2)alpha, (3) beta subunit tertiary structure changes that are located between residues Asp99(G1)beta and Asn102(G4)beta, (4) increased mobility of the alpha subunit COOH-terminal dipeptide, and (5) shortening of the Fe-Nepsilon2His(F8) bond in the alpha and beta subunits of the betaW37G and betaW37E mutants. In each case, the magnitude of the change in a particular structural parameter increases in the order betaW37Y < betaW37A < betaW37E approximately betaW37G, which corresponds closely to the degree of functional disruption documented in the preceding papers.
- Department of Biochemistry, College of Medicine, The University of Iowa, Iowa City, Iowa 52242, USA.
Organizational Affiliation: