Structure of human CCR4 in complex with AZD2098
Qi, X., Zhou, N., Wang, R.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| C-C chemokine receptor type 4,Soluble cytochrome b562 | A [auth R] | 475 | Homo sapiens, Escherichia coli | Mutation(s): 0 Gene Names: CCR4, CMKBR4, cybC | 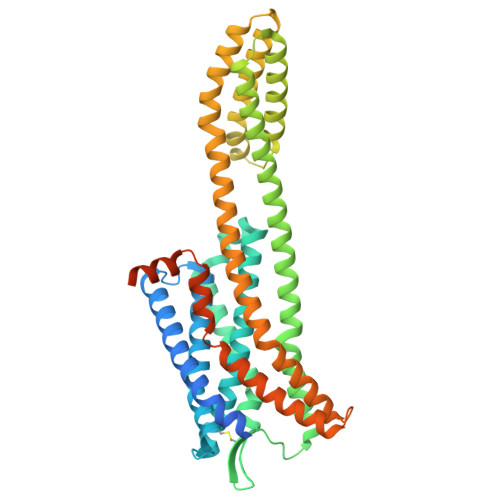 |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for P51679 (Homo sapiens) Explore P51679 Go to UniProtKB: P51679 | |||||
PHAROS: P51679 GTEx: ENSG00000183813 | |||||
Find proteins for P0ABE7 (Escherichia coli) Explore P0ABE7 Go to UniProtKB: P0ABE7 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Groups | P51679P0ABE7 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Anti-BRIL Fab heavy chain | B [auth H] | 229 | Homo sapiens | Mutation(s): 0 | 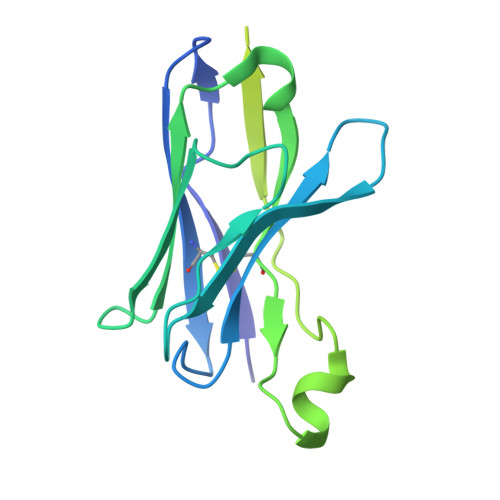 |
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Anti-BRIL Fab light chain | C [auth L] | 214 | Homo sapiens | Mutation(s): 0 | 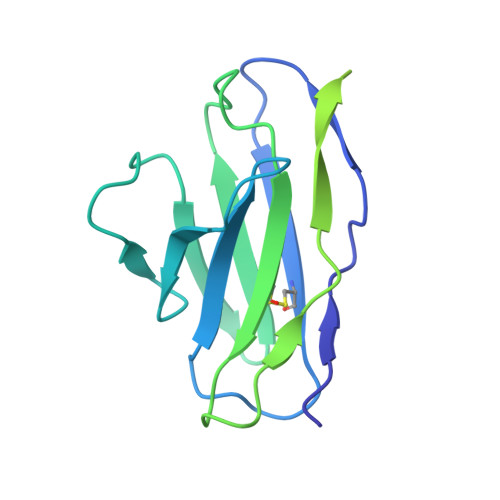 |
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| A1DAN (Subject of Investigation/LOI) Query on A1DAN | D [auth R] | 2,3-dichloro-N-(3-methoxypyrazin-2-yl)benzene-1-sulfonamide C11 H9 Cl2 N3 O3 S FLSMVCMSUNISFK-UHFFFAOYSA-N |  | ||
| Task | Software Package | Version |
|---|---|---|
| MODEL REFINEMENT | PHENIX | |
| RECONSTRUCTION | cryoSPARC |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Cancer Prevention and Research Institute of Texas (CPRIT) | United States | RR230054 |
| Welch Foundation | United States | I-2253-20250403 |