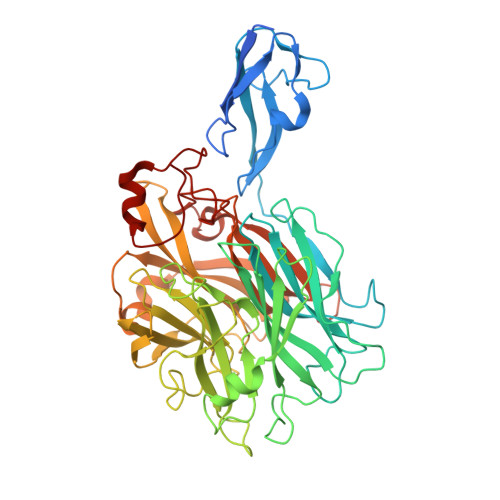
Bacterial exo-alpha-sialidases subvert the complement system through desialylation.
Kurniyati, K., Clark, N.D., Fu, Q., Zhang, S., Qiu, W., Malkowski, M.G., Li, C.(2026) bioRxiv
- PubMed: 41676663 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.64898/2026.02.05.703967
- Primary Citation Related Structures:
10QF, 10QG, 10QH, 10QI - PubMed Abstract:
The complement system is a central pillar of innate immunity, yet how bacterial glycan-modifying enzymes subvert complement function remains poorly understood. Bacterial sialidases remove terminal sialic acids from host sialoglycans, but their direct impact on complement immunity is unclear. Here, we investigate the impact of six sialidases from five human pathogens on complement immunity using an integrated approach combining genetics, biochemistry, glycobiology, mass spectrometry, and structural biology. We demonstrate that major complement components (IgG, C1q, C4, and C5) and regulators (Factor I, Factor H, and C4bp) are sialylated, and that bacterial sialidase-mediated desialylation suppresses complement activation and surface deposition, thereby enabling complement evasion. Despite extensive sequence diversity, biochemical and structural analyses reveal that all examined sialidases share a conserved six-bladed β-propeller catalytic domain and cleave both N-acetylneuraminic and N-glycolylneuraminic acids, the predominant mammalian sialic acids. Together, these findings uncover a conserved mechanism by which diverse bacterial pathogens disable complement immunity through desialylation.
- Philips Institute for Oral Health Research, Virginia Commonwealth University, Richmond, VA, USA.
Organizational Affiliation: