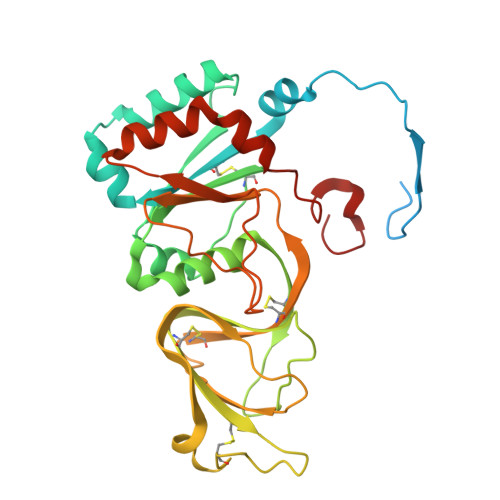
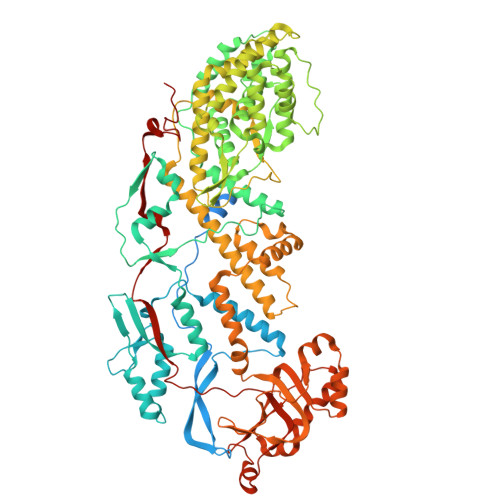
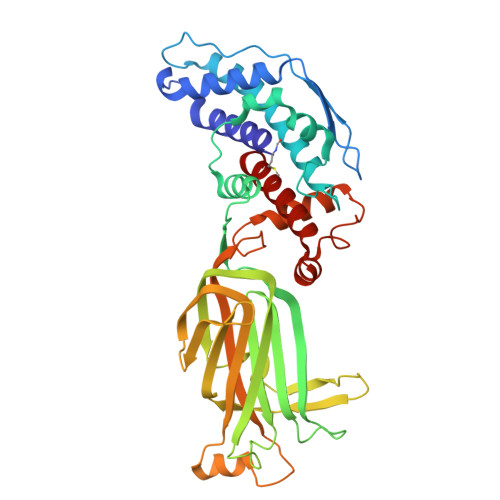
Mechanism of membrane perforation in rotavirus cell entry.
de Sautu, M., Leistner, C., Kirchhausen, T., Jenni, S., Harrison, S.C.(2026) bioRxiv
- PubMed: 41659395 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.64898/2026.01.21.700916
- Primary Citation Related Structures:
10IC, 10ID - PubMed Abstract:
Infectious cell entry by non-enveloped viruses requires delivery of the viral genome -- in many cases enclosed within a large, subviral particle -- across the membrane of an intracellular compartment. Rotaviruses and other double-strand RNA (dsRNA) viruses introduce into their target cells an inner capsid particle, roughly 700 Å in diameter, that does not uncoat further but instead extrudes capped viral mRNA by virtue of RNA-dependent RNA polymerase and capping activities within it. The delivery agent is an outer protein layer of the virion. We describe here use of cryogenic electron tomography (cryo-ET) to visualize the full course of rhesus rotavirus (RRV) entry, from cell attachment and inward budding of the virion to arrival of the subviral particle in the cytosol. The cryo-tomograms and subtomogram averaging of classified subparticles have enabled us to link high-resolution structures of the virion and its components with time series from live-cell fluorescence microscopy and thus to outline the molecular mechanism of each step in the entry process, including the hitherto elusive membrane perforation step needed for transfer of the subviral particle into the cytosol.