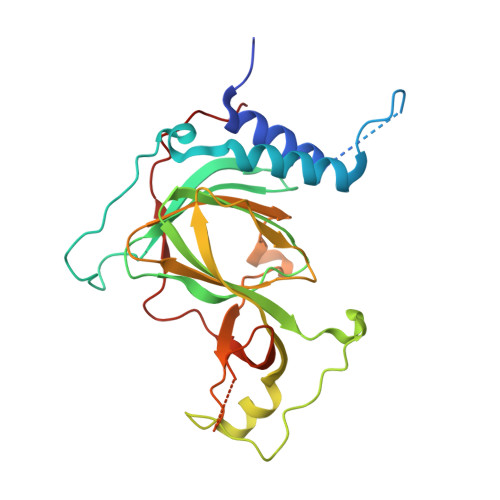
Cobalt(II)-Substituted Cysteamine Dioxygenase Oxygenation Proceeds through a Cobalt(III)-Superoxo Complex.
Li, J., Duan, R., Liu, A.(2024) J Am Chem Soc 146: 18292-18297
- PubMed: 38941563
- DOI: https://doi.org/10.1021/jacs.4c01871
- Primary Citation of Related Structures:
8U9J, 8UAN - PubMed Abstract:
We investigated the metal-substituted catalytic activity of human cysteamine dioxygenase (ADO), an enzyme pivotal in regulating thiol metabolism and contributing to oxygen homeostasis. Our findings demonstrate the catalytic competence of cobalt(II)- and nickel(II)-substituted ADO in cysteamine oxygenation. Notably, Co(II)-ADO exhibited superiority over Ni(II)-ADO despite remaining significantly less active than the natural enzyme. Structural analyses through X-ray crystallography and cobalt K-edge excitation confirmed successful metal substitution with minimal structural perturbations. This provided a robust structural basis, supporting a conserved catalytic mechanism tailored to distinct metal centers. This finding challenges the proposed high-valent ferryl-based mechanism for thiol dioxygenases, suggesting a non-high-valent catalytic pathway in the native enzyme. Further investigation of the cysteamine-bound or a peptide mimic of N -terminus RGS5 bound Co(II)-ADO binary complex revealed the metal center's high-spin ( S = 3/2) state. Upon reaction with O 2 , a kinetically and spectroscopically detectable intermediate emerged with a ground spin state of S = 1/2. This intermediate exhibits a characteristic 59 Co hyperfine splitting ( A = 67 MHz) structure in the EPR spectrum alongside UV-vis features, consistent with known low-spin Co(III)-superoxo complexes. This observation, unique for protein-bound thiolate-ligated cobalt centers in a protein, unveils the capacities for O 2 activation in such metal environments. These findings provide valuable insights into the non-heme iron-dependent thiol dioxygenase mechanistic landscape, furthering our understanding of thiol metabolism regulation. The exploration of metal-substituted ADO sheds light on the intricate interplay between metal and catalytic activity in this essential enzyme.
Organizational Affiliation:
Department of Chemistry, University of Texas at San Antonio, San Antonio, Texas 78249, United States.