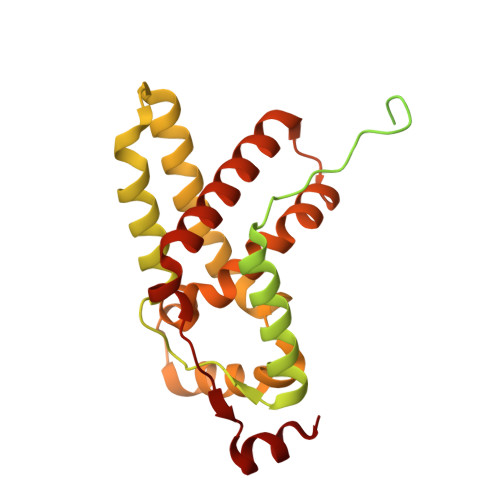
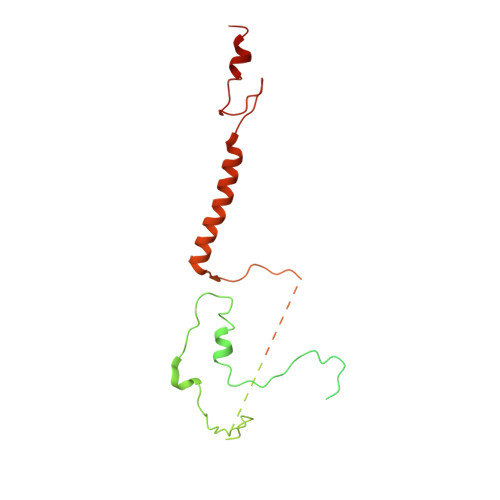
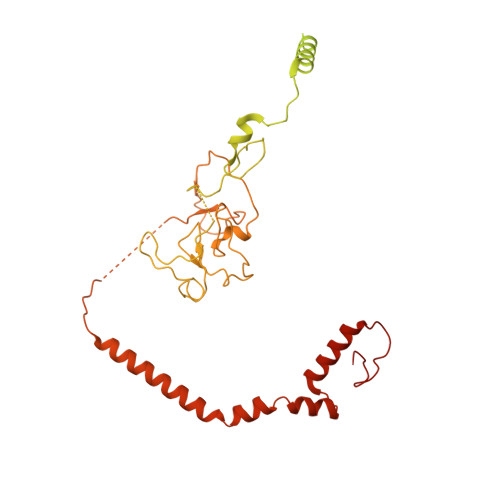
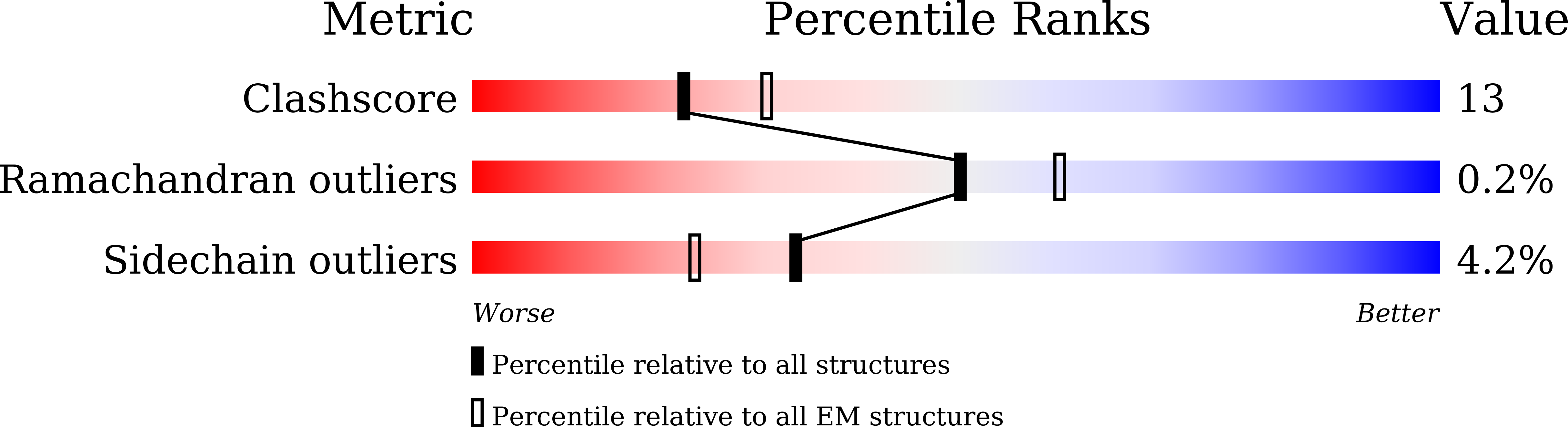Class I histone deacetylase complex: Structure and functional correlates.
Wang, X., Wang, Y., Liu, S., Zhang, Y., Xu, K., Ji, L., Kornberg, R.D., Zhang, H.(2023) Proc Natl Acad Sci U S A 120: e2307598120
- PubMed: 37459529
- DOI: https://doi.org/10.1073/pnas.2307598120
- Primary Citation of Related Structures:
8IFG - PubMed Abstract:
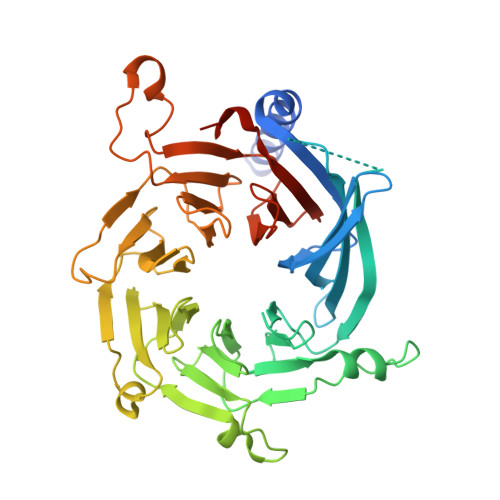
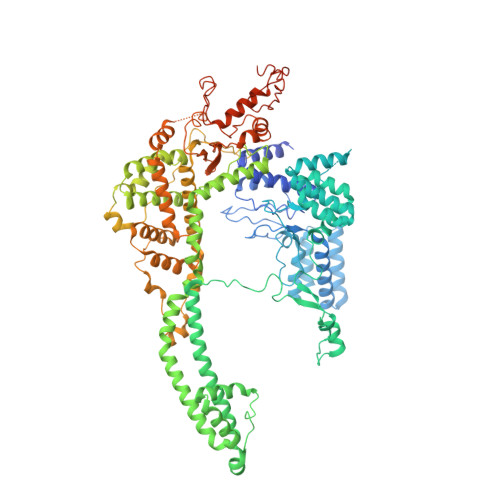
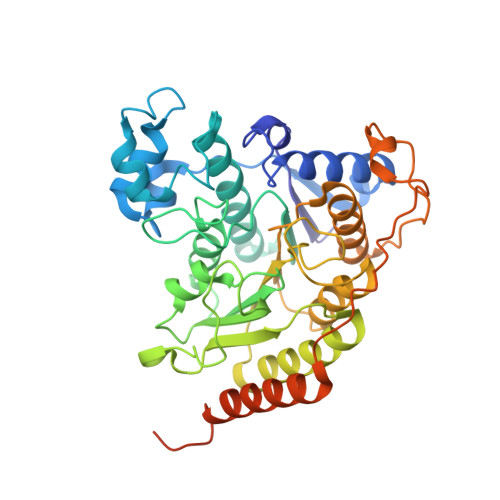
The Schizosaccharomyces pombe Clr6S complex, a class I histone deacetylase complex, functions as a zinc-dependent enzyme to remove acetyl groups from lysine residues in histone tails. We report here the cryo-EM structure of Clr6S alone and a cryo-EM map of Clr6S in complex with a nucleosome. The active center, revealed at near-atomic resolution, includes features important for catalysis-A water molecule coordinated by zinc, the likely nucleophile for attack on the acetyl-lysine bond, and a loop that may position the substrate for catalysis. The cryo-EM map in the presence of a nucleosome reveals multiple Clr6S-nucleosome contacts and a high degree of relative motion of Clr6S and the nucleosome. Such flexibility may be attributed to interaction at a site in the flexible histone tail and is likely important for the function of the deacetylase, which acts at multiple sites in other histone tails.
Organizational Affiliation:
Shanghai Institute for Advanced Immunochemical Studies, ShanghaiTech University, Shanghai 201210, China.