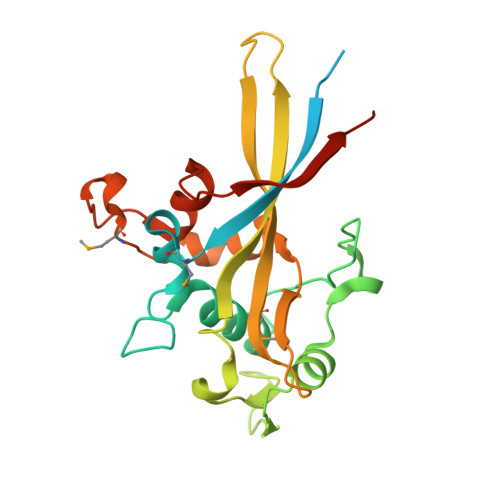
Structural and functional analyses of the Porphyromonas gingivalis type IX secretion system PorN protein.
Fuchsbauer, O., Lunar Silva, I., Cascales, E., Roussel, A., Leone, P.(2022) J Biol Chem 298: 101618-101618
- PubMed: 35065963
- DOI: https://doi.org/10.1016/j.jbc.2022.101618
- Primary Citation of Related Structures:
7PVH - PubMed Abstract:
Porphyromonas gingivalis, the major human pathogen bacterium associated with periodontal diseases, secretes virulence factors through the Bacteroidetes-specific type IX secretion system (T9SS). Effector proteins of the T9SS are recognized by the complex via their conserved C-terminal domains (CTDs). Among the 18 proteins essential for T9SS function in P. gingivalis, PorN is a periplasmic protein that forms large ring-shaped structures in association with the PorK outer membrane lipoprotein. PorN also mediates contacts with the PorM subunit of the PorLM energetic module, and with the effector's CTD. However, no information is available on the PorN structure and on the implication of PorN domains for T9SS assembly and effector recognition. Here we present the crystal structure of PorN at 2.0-Å resolution, which represents a novel fold with no significant similarity to any known structure. In agreement with in silico analyses, we also found that the N- and C-terminal regions of PorN are intrinsically disordered. Our functional studies showed that the N-terminal disordered region is involved in PorN dimerization while the C-terminal disordered region is involved in the interaction with PorK. Finally, we determined that the folded PorN central domain is involved in the interaction with PorM, as well as with the effector's CTD. Altogether, these results lay the foundations for a more comprehensive model of T9SS architecture and effector transport.
Organizational Affiliation:
Architecture et Fonction des Macromolécules Biologiques, Aix-Marseille Université, UMR 7257, Marseille, France; Architecture et Fonction des Macromolécules Biologiques, Centre National de la Recherche Scientifique, UMR 7257, Marseille, France.