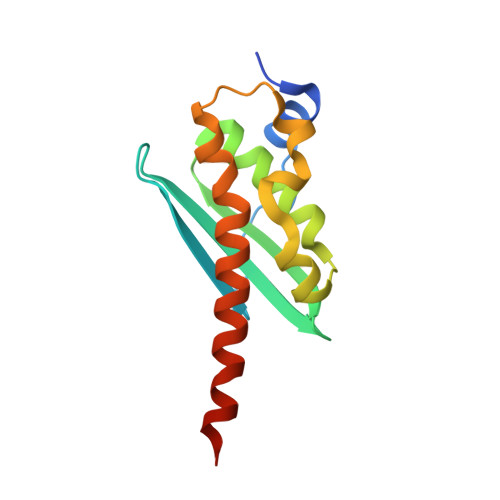
The Structure of the Dimeric State of IscU Harboring Two Adjacent [2Fe-2S] Clusters Provides Mechanistic Insights into Cluster Conversion to [4Fe-4S].
Kunichika, K., Nakamura, R., Fujishiro, T., Takahashi, Y.(2021) Biochemistry 60: 1569-1572
- PubMed: 33938220
- DOI: https://doi.org/10.1021/acs.biochem.1c00112
- Primary Citation of Related Structures:
7C8M, 7C8N - PubMed Abstract:
IscU serves as a scaffold for the de novo assembly of a [2Fe-2S] cluster prior to its delivery to recipient protein. It has also been proposed that on one dimer of bacterial IscU, two [2Fe-2S] clusters can be converted into a single [4Fe-4S] cluster. However, lack of structural information about the dimeric state of IscU has hindered our understanding of the underlying mechanisms. In this study, we determine the X-ray crystal structure of IscU from the thermophilic archaeon Methanothrix thermoacetophila and demonstrate a dimer structure of IscU in which two [2Fe-2S] clusters are facing each other in close proximity at the dimer interface. Our structure also reveals for the first time that Asp40 serves as a fourth ligand to the [2Fe-2S] cluster with three Cys ligands in each monomer, consistent with previous spectroscopic data. We confirm by EPR spectroscopic analysis that in solution two adjacent [2Fe-2S] clusters in the wild-type dimer are converted to a [4Fe-4S] cluster via reductive coupling. Furthermore, we find that the H106A substitution abolishes the reductive conversion to the [4Fe-4S] cluster without structural alteration, suggesting that His106 is functionally involved in this process. Overall, these findings provide a structural explanation for the assembly and conversion of Fe-S clusters on IscU and highlight a dynamic process that advances via association and dissociation of the IscU dimer.
Organizational Affiliation:
Department of Biochemistry and Molecular Biology, Graduate School of Science and Engineering, Saitama University, Shimo-okubo 255, Sakura-ku, Saitama 338-8570, Japan.