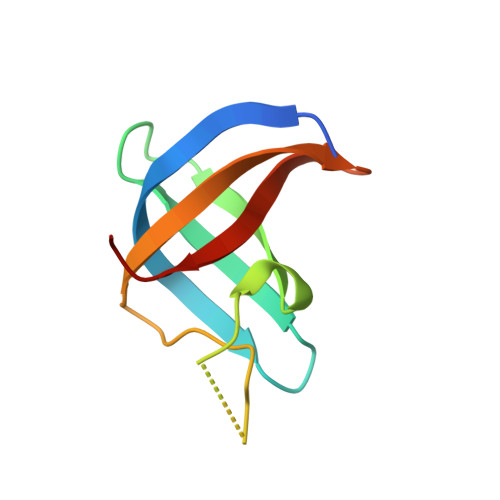
Crystal structure of a Y-box binding protein 1 (YB-1)-RNA complex reveals key features and residues interacting with RNA.
Yang, X.J., Zhu, H., Mu, S.R., Wei, W.J., Yuan, X., Wang, M., Liu, Y., Hui, J., Huang, Y.(2019) J Biol Chem 294: 10998-11010
- PubMed: 31160337
- DOI: https://doi.org/10.1074/jbc.RA119.007545
- Primary Citation of Related Structures:
5YTS, 5YTT, 5YTV, 5YTX - PubMed Abstract:
The Y-box binding protein 1 (YB-1) is a member of the cold shock domain (CSD) protein family and is recognized as an oncogenic factor in several solid tumors. By binding to RNA, YB-1 participates in several steps of posttranscriptional regulation of gene expression, including mRNA splicing, stability, and translation; microRNA processing; and stress granule assembly. However, the mechanisms in YB-1-mediated regulation of RNAs are unclear. Previously, we used both systematic evolution of ligands by exponential enrichment (SELEX) and individual-nucleotide resolution UV cross-linking and immunoprecipitation coupled RNA-Seq (iCLIP-Seq) analyses, which defined the RNA-binding consensus sequence of YB-1 as CA(U/C)C. We also reported that through binding to its core motif CAUC in primary transcripts, YB-1 regulates the alternative splicing of a CD44 variable exon and the biogenesis of miR-29b-2 during both Drosha and Dicer steps. To elucidate the molecular basis of the YB-1-RNA interactions, we report high-resolution crystal structures of the YB-1 CSD in complex with different RNA oligos at 1.7 Å resolution. The structure revealed that CSD interacts with RNA mainly through π-π stacking interactions assembled by four highly conserved aromatic residues. Interestingly, YB-1 CSD forms a homodimer in solution, and we observed that two residues, Tyr-99 and Asp-105, at the dimer interface are important for YB-1 CSD dimerization. Substituting these two residues with Ala reduced CSD's RNA-binding activity and abrogated the splicing activation of YB-1 targets. The YB-1 CSD-RNA structures presented here at atomic resolution provide mechanistic insights into gene expression regulated by CSD-containing proteins.
Organizational Affiliation:
CAS Center for Excellence in Molecular Cell Science, Shanghai 200031, China,; State Key Laboratory of Molecular Biology, Shanghai Institute of Biochemistry and Cell Biology, Chinese Academy of Sciences, University of Chinese Academy of Sciences, Shanghai 200031, China, and; Shanghai Key Laboratory of Molecular Andrology, Shanghai 200031, China.