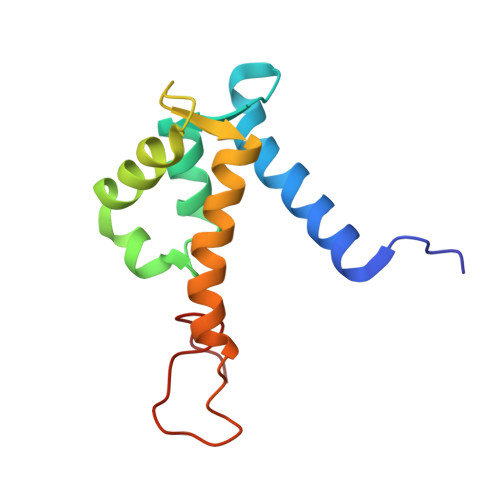
Blocking the interaction between S100A9 and RAGE V domain using CHAPS molecule: A novel route to drug development against cell proliferation
Chang, C.-C., Khan, I., Tsai, K.-L., Li, H., Yang, L.-W., Chou, R.-H., Yu, C.(2016) Biochim Biophys Acta 1864: 1558-1569
- PubMed: 27524699
- DOI: https://doi.org/10.1016/j.bbapap.2016.08.008
- Primary Citation of Related Structures:
5I8N - PubMed Abstract:
Human S100A9 (Calgranulin B) is a Ca(2+)-binding protein, from the S100 family, that often presents as a homodimer in myeloid cells. It becomes an important mediator during inflammation once calcium binds to its EF-hand motifs. Human RAGE protein (receptor for advanced glycation end products) is one of the target-proteins. RAGE binds to a hydrophobic surface on S100A9. Interactions between these proteins trigger signal transduction cascades, promoting cell growth, proliferation, and tumorigenesis. Here, we present the solution structure of mutant S100A9 (C3S) homodimer, determined by multi-dimensional NMR experiments. We further characterize the solution interactions between mS100A9 and the RAGE V domain via NMR spectroscopy. CHAPS is a zwitterionic and non-denaturing molecule widely used for protein solubilizing and stabilization. We found out that CHAPS and RAGE V domain would interact with mS100A9 by using (1)H-(15)N HSQC NMR titrations. Therefore, using the HADDOCK program, we superimpose two binary complex models mS100A9-RAGE V domain and mS100A9-CHAPS and demonstrate that CHAPS molecules could play a crucial role in blocking the interaction between mS100A9 and the RAGE V domain. WST-1 assay results also support the conclusion that CHAPS inhibits the bioactivity of mS100A9. This report will help to inform new drug development against cell proliferation.
Organizational Affiliation:
Department of Chemistry, National Tsing Hua University, Hsinchu 30013, Taiwan.