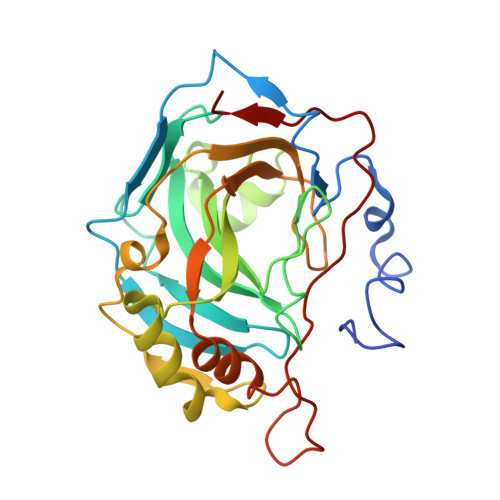
Carbonic anhydrase inhibitors. X-ray crystal studies of the carbonic anhydrase II-trithiocarbonate adduct-An inhibitor mimicking the sulfonamide and urea binding to the enzyme
Temperini, C., Scozzafava, A., Supuran, C.T.(2010) Bioorg Med Chem Lett 20: 474-478
- PubMed: 20005709
- DOI: https://doi.org/10.1016/j.bmcl.2009.11.124
- Primary Citation of Related Structures:
3K7K - PubMed Abstract:
Trithiocarbonate (CS32-) inhibits with low micromolar affinities several mammalian carbonic anhydrases, CAs, EC 4.2.1.1 [Innocenti et al., Bioorg. Med. Chem. Lett. 2009, 19, 1855]. Here we report the X-ray crystal structure of the hCA II-trithiocarbonate adduct. Trithiocarbonate is monodentately bound to the Zn(II) ion and makes several hydrogen bonds with Thr199 and two water molecules from the enzyme active site. Its binding is different from that of ureate, another small inhibitor isosteric with trithiocarbonate but somehow mimicks the binding of the SO(2)NH moiety present in the sulfonamide inhibitors and is similar to that of bicarbonate. Compounds incorporating this new zinc-binding group, CS2-, may thus lead to new classes of potent inhibitors.
Organizational Affiliation:
Università degli Studi di Firenze, Laboratorio di Chimica Bioinorganica, Rm. 188, Via della Lastruccia 3, I-50019 Sesto Fiorentino (Firenze), Italy.