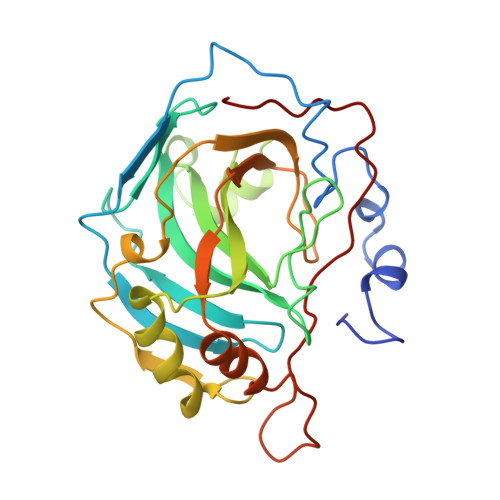
Unexpected binding mode of the sulfonamide fluorophore 5-dimethylamino-1-naphthalene sulfonamide to human carbonic anhydrase II. Implications for the development of a zinc biosensor.
Nair, S.K., Elbaum, D., Christianson, D.W.(1996) J Biol Chem 271: 1003-1007
- PubMed: 8557623
- DOI: https://doi.org/10.1074/jbc.271.2.1003
- Primary Citation of Related Structures:
1OKL - PubMed Abstract:
The three-dimensional structure of human carbonic anhydrase II (CAII) complexed with the sulfonamide fluorophore 5-dimethylamino-1-naphthalene sulfonamide (dansylamide) has been determined to 2.1-A resolution by x-ray crystallographic methods. Unlike other arylsulfonamide inhibitors of CAII, the naphthyl ring of dansylamide binds in a hydrophobic pocket in the active site, making van der Waals contacts with Val-121, Phe-131, Val-143, Leu-198, and Trp-209. Interestingly, a conformational change of Leu-198 is required to accommodate dansylamide binding, which rationalizes the enhanced dansylamide affinity measured for certain Leu-198 variants (Nair, S. K., Krebs, J.F., Christianson, D. W., and Fierke, C. A. (1995) Biochemistry 34, 3981-3989). Modeling studies indicate that a second binding mode, in which the fused aromatic ring is rotated out of the hydrophobic pocket, is sterically feasible. Both experimentally observed and modeled binding modes have implications for new leads in the design of avid CAII inhibitors. Finally, the structure of the CAII-dansylamide complex has implications for its exploitation in zinc biosensor applications, and possible routes toward the optimization of fluorophore design are considered on the basis on this structure.
Organizational Affiliation:
Department of Chemistry, University of Pennsylvania, Philadelphia 19104-6323, USA.