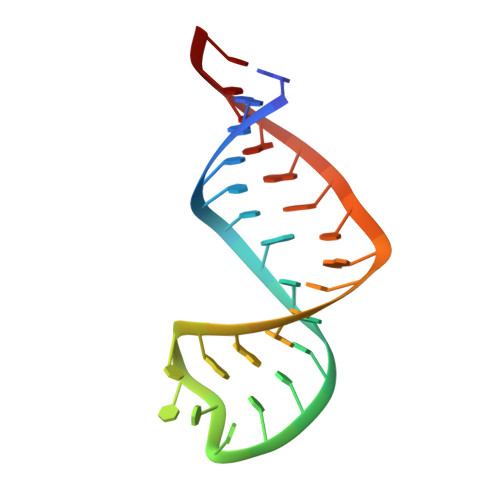
The 5'-terminal stem-loop RNA element of SARS-CoV-2 features highly dynamic structural elements that are sensitive to differences in cellular pH.
Toews, S., Wacker, A., Faison, E.M., Duchardt-Ferner, E., Richter, C., Mathieu, D., Bottaro, S., Zhang, Q., Schwalbe, H.(2024) Nucleic Acids Res 52: 7971-7986
- PubMed: 38842942
- DOI: https://doi.org/10.1093/nar/gkae477
- Primary Citation of Related Structures:
9EOW - PubMed Abstract:
We present the nuclear magnetic resonance spectroscopy (NMR) solution structure of the 5'-terminal stem loop 5_SL1 (SL1) of the SARS-CoV-2 genome. SL1 contains two A-form helical elements and two regions with non-canonical structure, namely an apical pyrimidine-rich loop and an asymmetric internal loop with one and two nucleotides at the 5'- and 3'-terminal part of the sequence, respectively. The conformational ensemble representing the averaged solution structure of SL1 was validated using NMR residual dipolar coupling (RDC) and small-angle X-ray scattering (SAXS) data. We show that the internal loop is the major binding site for fragments of low molecular weight. This internal loop of SL1 can be stabilized by an A12-C28 interaction that promotes the transient formation of an A+•C base pair. As a consequence, the pKa of the internal loop adenosine A12 is shifted to 5.8, compared to a pKa of 3.63 of free adenosine. Furthermore, applying a recently developed pH-differential mutational profiling (PD-MaP) approach, we not only recapitulated our NMR findings of SL1 but also unveiled multiple sites potentially sensitive to pH across the 5'-UTR of SARS-CoV-2.
Organizational Affiliation:
Institute of Organic Chemistry and Chemical Biology, Johann Wolfgang Goethe-University Frankfurt, Frankfurt/Main, Hesse 60438, Germany.