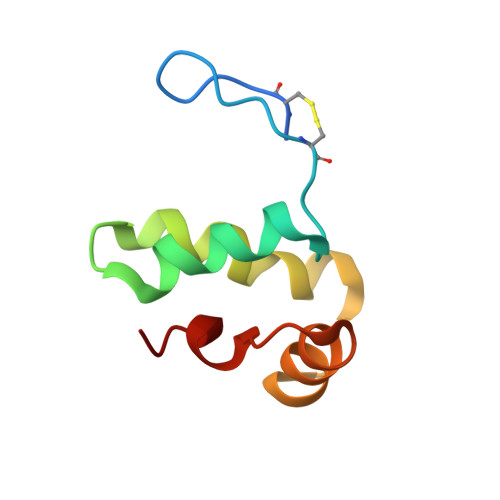
Staphylococcal Peroxidase Inhibitor (SPIN): Investigation of the Inhibitory N-terminal Domain via a Stabilizing Disulfide Insertion.
Fatehi, S., Mishra, N., Herdendorf, T.J., Prakash, O., Geisbrecht, B.V.(2024) Arch Biochem Biophys : 110060-110060
- PubMed: 38880318
- DOI: https://doi.org/10.1016/j.abb.2024.110060
- Primary Citation of Related Structures:
9BD0 - PubMed Abstract:
Staphylococcus aureus secretes an array of small proteins that inhibit key enzyme-catalyzed reactions necessary for proper function of the human innate immune system. Among these, the Staphylococcal Peroxidase Inhibitor, SPIN, blocks the activity of myeloperoxidase (MPO) and thereby disrupts the HOCl-generating system of neutrophils. Previous studies on S. aureus SPIN have shown that it relies on a C-terminal α-helical bundle domain to mediate initial binding to MPO, but requires a disordered N-terminal region to fold into a β-hairpin conformation to inhibit MPO activity. To further investigate the structure/function relationship of SPIN, we introduced two cysteine residues into its N-terminal region to trap SPIN in its MPO-bound conformation and characterized the modified protein, which we refer to here as SPIN-CYS. Although control experiments confirmed the presence of the disulfide bond in SPIN-CYS, solution structure determination revealed that the N-terminal region of SPIN-CYS adopted a physically constrained series of lariat-like structures rather than a well-defined β-hairpin. Nevertheless, SPIN-CYS exhibited a gain in inhibitory potency against human MPO when compared to wild-type SPIN. This gain of function persisted even in the presence of deleterious mutations within the C-terminal α-helical bundle domain. Surface plasmon resonance studies showed that the gain in potency arose through an increase in apparent affinity of SPIN-CYS for MPO, which was driven primarily by an increased association rate with MPO when compared to wild-type SPIN. Together, this work provides new information on the coupled binding and folding events required to manifest biological activity of this unusual MPO inhibitor.
Organizational Affiliation:
Department of Biochemistry & Molecular Biophysics, Kansas State University, Manhattan, KS, 66506, United States.