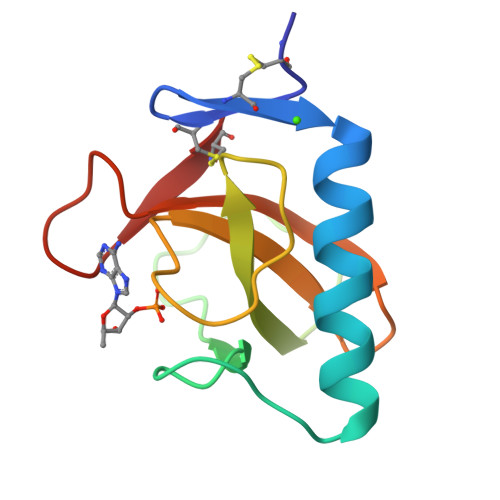
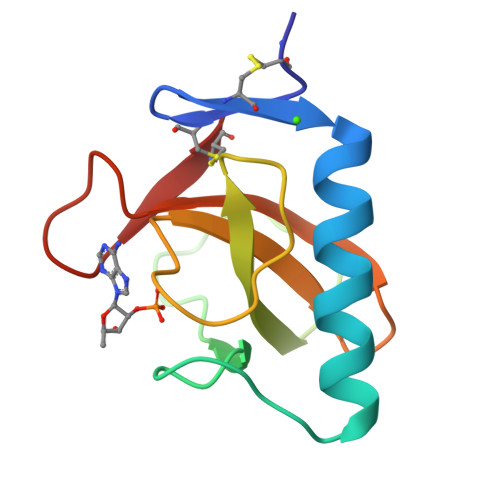
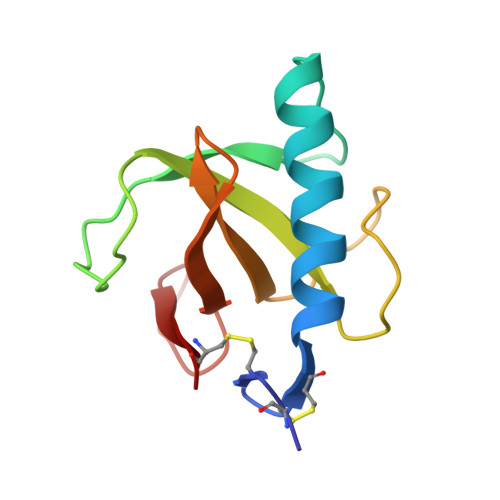
Crystal structure of the Tyr45Trp mutant of ribonuclease T1 in a complex with 2'-adenylic acid.
Koellner, G., Grunert, H.P., Landt, O., Saenger, W.(1991) Eur J Biochem 201: 199-202
- PubMed: 1915364
- DOI: https://doi.org/10.1111/j.1432-1033.1991.tb16274.x
- Primary Citation of Related Structures:
7RNT - PubMed Abstract:
The recombinant Tyr45Trp mutant of Lys25-ribonuclease T1 was overexpressed and purified from an Escherichia coli strain. The mutant enzyme, which shows reduced activity towards GpA and increased activity towards pGpC, pApC and pUpC compared with wild-type RNase T1, was co-crystallized with 2'-adenylic acid by microdialysis. The space group is P212121 with unit cell dimenions a = 4.932(2), b = 4.661(2), c = 4.092(1) nm. The crystal structure was solved using the coordinates of the isomorphous complex of wild-type RNase T1 with 2'-AMP. The refinement was based on Fhkl of 7726 reflexions with Fo greater than or equal to 1 sigma (Fo) in the resolution range of 2.0-0.19 nm and converged with an R factor of 0.179. The adenosine of 2'-AMP is not bound to the guanosine binding site, as could be expected from the mutation of Tyr45Trp, but is stacked on the Gly74 carbonyl group and the His92 imidazole group which form a subsite for substrate binding, as already observed in the wild-type 2'-AMP complex. The point mutation of Tyr45Trp does not perturb the backbone conformation and the Trp-indole side chain is in a comparable position to the phenolic Tyr45 of the wild-type enzyme.
Organizational Affiliation:
Institut für Kristallographie, Freie Universität Berlin, Federal Republic of Germany.