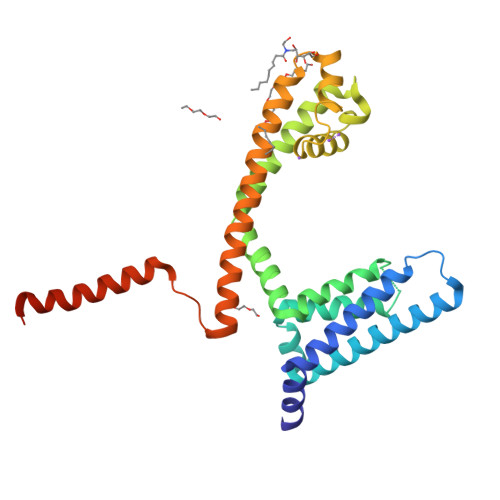
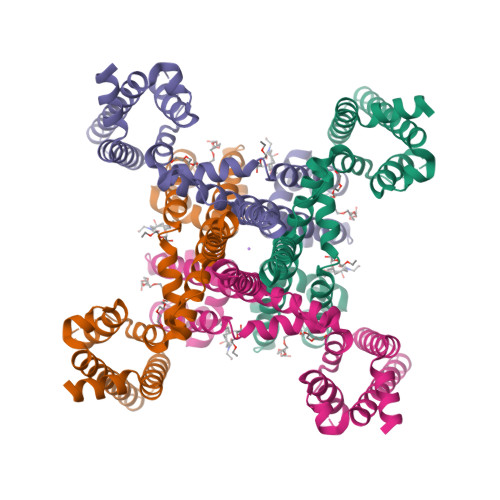
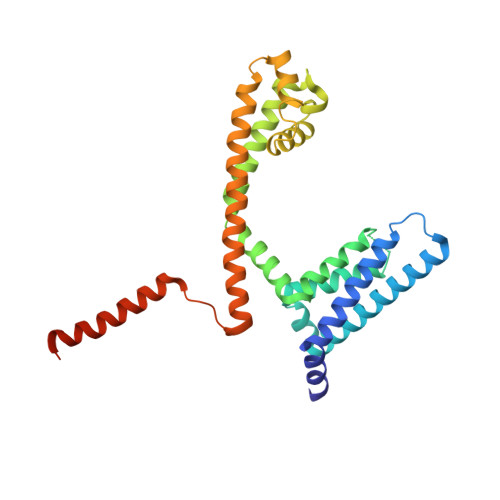
Cannabidiol interactions with voltage-gated sodium channels.
Sait, L.G., Sula, A., Ghovanloo, M.R., Hollingworth, D., Ruben, P.C., Wallace, B.A.(2020) Elife 9
- PubMed: 33089780
- DOI: https://doi.org/10.7554/eLife.58593
- Primary Citation of Related Structures:
6YZ0, 6YZ2 - PubMed Abstract:
Voltage-gated sodium channels are targets for a range of pharmaceutical drugs developed for the treatment of neurological diseases. Cannabidiol (CBD), the non-psychoactive compound isolated from cannabis plants, was recently approved for treatment of two types of epilepsy associated with sodium channel mutations. This study used high-resolution X-ray crystallography to demonstrate the detailed nature of the interactions between CBD and the NavMs voltage-gated sodium channel, and electrophysiology to show the functional effects of binding CBD to these channels. CBD binds at a novel site at the interface of the fenestrations and the central hydrophobic cavity of the channel. Binding at this site blocks the transmembrane-spanning sodium ion translocation pathway, providing a molecular mechanism for channel inhibition. Modelling studies suggest why the closely-related psychoactive compound tetrahydrocannabinol may not have the same effects on these channels. Finally, comparisons are made with the TRPV2 channel, also recently proposed as a target site for CBD. In summary, this study provides novel insight into a possible mechanism for CBD interactions with sodium channels.
Organizational Affiliation:
Institute of Structural and Molecular Biology, Birkbeck College, University of London, London, United Kingdom.