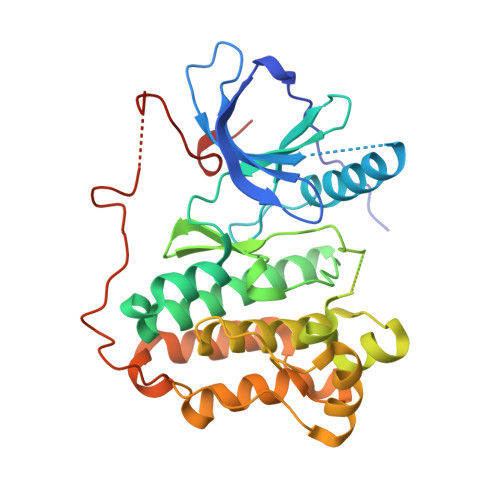
Discovery of Selective and Noncovalent Diaminopyrimidine-Based Inhibitors of Epidermal Growth Factor Receptor Containing the T790M Resistance Mutation.
Hanan, E.J., Eigenbrot, C., Bryan, M.C., Burdick, D.J., Chan, B.K., Chen, Y., Dotson, J., Heald, R.A., Jackson, P.S., La, H., Lainchbury, M.D., Malek, S., Purkey, H.E., Schaefer, G., Schmidt, S., Seward, E.M., Sideris, S., Tam, C., Wang, S., Yeap, S.K., Yen, I., Yin, J., Yu, C., Zilberleyb, I., Heffron, T.P.(2014) J Med Chem 57: 10176-10191
- PubMed: 25383627
- DOI: https://doi.org/10.1021/jm501578n
- Primary Citation of Related Structures:
4RJ3, 4RJ4, 4RJ5, 4RJ6, 4RJ7, 4RJ8 - PubMed Abstract:
Activating mutations within the epidermal growth factor receptor (EGFR) kinase domain, commonly L858R or deletions within exon 19, increase EGFR-driven cell proliferation and survival and are correlated with impressive responses to the EGFR inhibitors erlotinib and gefitinib in nonsmall cell lung cancer patients. Approximately 60% of acquired resistance to these agents is driven by a single secondary mutation within the EGFR kinase domain, specifically substitution of the gatekeeper residue threonine-790 with methionine (T790M). Due to dose-limiting toxicities associated with inhibition of wild-type EGFR (wtEGFR), we sought inhibitors of T790M-containing EGFR mutants with selectivity over wtEGFR. We describe the evolution of HTS hits derived from Jak2/Tyk2 inhibitors into selective EGFR inhibitors. X-ray crystal structures revealed two distinct binding modes and enabled the design of a selective series of novel diaminopyrimidine-based inhibitors with good potency against T790M-containing mutants of EGFR, high selectivity over wtEGFR, broad kinase selectivity, and desirable physicochemical properties.
Organizational Affiliation:
Departments of †Discovery Chemistry, ‡Structural Biology, §Drug Metabolism and Pharmacokinetics, ∥Biochemical and Cellular Pharmacology, ⊥Molecular Oncology, and #Protein Expression, Genentech Inc. , 1 DNA Way, South San Francisco, California 94080, United States.