Conformationally Defined Rexinoids and Their Efficacy in the Prevention of Mammary Cancers.
Atigadda, V.R., Xia, G., Deshpande, A., Wu, L., Kedishvili, N., Smith, C.D., Krontiras, H., Bland, K.I., Grubbs, C.J., Brouillette, W.J., Muccio, D.D.(2015) J Med Chem 58: 7763-7774
- PubMed: 26331194
- DOI: https://doi.org/10.1021/acs.jmedchem.5b00829
- Primary Citation of Related Structures:
4RFW, 4RMC, 4RMD, 4RME - PubMed Abstract:
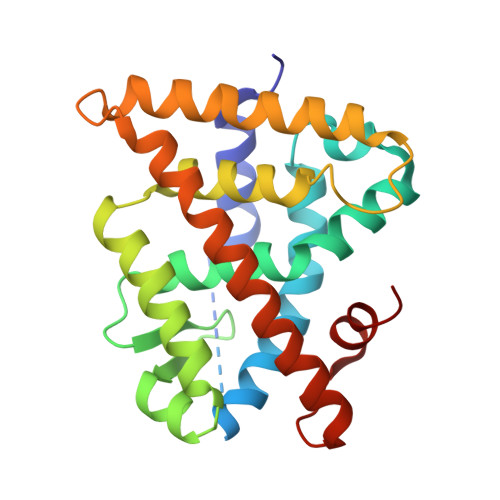
(2E,4E,6Z,8Z)-8-(3',4'-Dihydro-1'(2H)-naphthalen-1'-ylidene)-3,7-dimethyl-2,3,6-octatrienoinic acid (UAB30) is currently undergoing clinical evaluation as a novel cancer prevention agent. In efforts to develop even more highly potent rexinoids that prevent breast cancer without toxicity, we further explore here the structure-activity relationship of two separate classes of rexinoids. UAB30 belongs to the class II rexinoids and possesses a 9Z-tetraenoic acid chain bonded to a tetralone ring, whereas the class I rexinoids contain the same 9Z-tetraenoic acid chain bonded to a disubstituted cyclohexenyl ring. Among the 12 class I and class II rexinoids evaluated, the class I rexinoid 11 is most effective in preventing breast cancers in an in vivo rat model alone or in combination with tamoxifen. Rexinoid 11 also reduces the size of established tumors and exhibits a therapeutic effect. However, 11 induces hypertriglyceridemia at its effective dose. On the other hand rexinoid 10 does not increase triglyceride levels while being effective in the in vivo chemoprevention assay. X-ray studies of four rexinoids bound to the ligand binding domain of the retinoid X receptor reveal key structural aspects that enhance potency as well as those that enhance the synthesis of lipids.
Organizational Affiliation:
Departments of †Chemistry, ‡Biochemistry and Molecular Genetics, §Vision Sciences, and ∥Surgery, University of Alabama at Birmingham , Birmingham, Alabama 35294, United States.