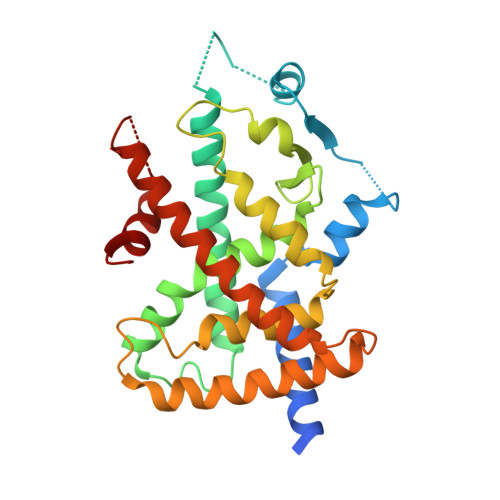
Structural basis of the transactivation deficiency of the human PPAR gamma F360L mutant associated with familial partial lipodystrophy.
Lori, C., Pasquo, A., Montanari, R., Capelli, D., Consalvi, V., Chiaraluce, R., Cervoni, L., Loiodice, F., Laghezza, A., Aschi, M., Giorgi, A., Pochetti, G.(2014) Acta Crystallogr D Biol Crystallogr 70: 1965-1976
- PubMed: 25004973
- DOI: https://doi.org/10.1107/S1399004714009638
- Primary Citation of Related Structures:
4L96, 4L98, 4O8F - PubMed Abstract:
The peroxisome proliferator-activated receptors (PPARs) are transcription factors that regulate glucose and lipid metabolism. The role of PPARs in several chronic diseases such as type 2 diabetes, obesity and atherosclerosis is well known and, for this reason, they are the targets of antidiabetic and hypolipidaemic drugs. In the last decade, some rare mutations in human PPARγ that might be associated with partial lipodystrophy, dyslipidaemia, insulin resistance and colon cancer have emerged. In particular, the F360L mutant of PPARγ (PPARγ2 residue 388), which is associated with familial partial lipodystrophy, significantly decreases basal transcriptional activity and impairs stimulation by synthetic ligands. To date, the structural reason for this defective behaviour is unclear. Therefore, the crystal structure of PPARγ F360L together with the partial agonist LT175 has been solved and the mutant has been characterized by circular-dichroism spectroscopy (CD) in order to compare its thermal stability with that of the wild-type receptor. The X-ray analysis showed that the mutation induces dramatic conformational changes in the C-terminal part of the receptor ligand-binding domain (LBD) owing to the loss of van der Waals interactions made by the Phe360 residue in the wild type and an important salt bridge made by Arg357, with consequent rearrangement of loop 11/12 and the activation function helix 12 (H12). The increased mobility of H12 makes the binding of co-activators in the hydrophobic cleft less efficient, thereby markedly lowering the transactivation activity. The spectroscopic analysis in solution and molecular-dynamics (MD) simulations provided results which were in agreement and consistent with the mutant conformational changes observed by X-ray analysis. Moreover, to evaluate the importance of the salt bridge made by Arg357, the crystal structure of the PPARγ R357A mutant in complex with the agonist rosiglitazone has been solved.
Organizational Affiliation:
Istituto di Cristallografia, CNR, Via Salaria, Km 29,300, Monterotondo Stazione, 00015 Roma, Italy.