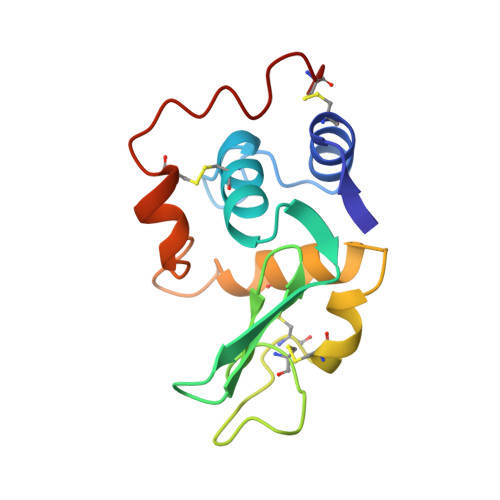
Crystal structures of the apo- and holomutant human lysozymes with an introduced Ca2+ binding site.
Inaka, K., Kuroki, R., Kikuchi, M., Matsushima, M.(1991) J Biol Chem 266: 20666-20671
- PubMed: 1939116
- Primary Citation of Related Structures:
2LHM, 3LHM - PubMed Abstract:
The three-dimensional structures of apo- and holomutant human lysozymes (D86/92 lysozyme), in which a calcium binding site was designed and created for enhancing molecular stability by replacing both Gln86 and Ala92 with aspartic acids, were refined at 1.8-A resolution by x-ray crystallography. The overall structures and crystallographic thermal factors of all three proteins, the apo-, holo-D86/92, and the wild-type human lysozymes, were essentially identical; these results showed that the introduction of the calcium binding site did not affect either the overall structure or molecular rigidity of the proteins. However, structure analyses of the apo-D86/92 lysozyme revealed that the mutations affected the side chain conformation of residue 86 and hydrogen networks between the protein and the internal solvent molecules. In the structure of the holo-D86/92 lysozyme, seven oxygen ligands formed a slightly distorted pentagonal bipyramid around the calcium ion, indicating that the coordination around the calcium ion was quite similar to that in baboon alpha-lactalbumin. The pentagonal bipyramid coordination could be one of the most widely found and appropriate calcium binding schemes in proteins.
Organizational Affiliation:
Protein Engineering Research Institute, Osaka, Japan.