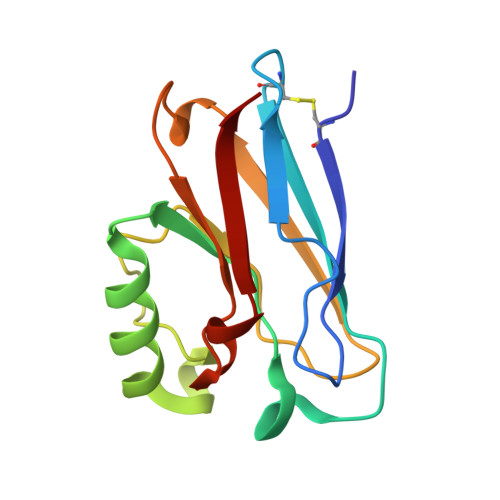
Relaxation dynamics of Pseudomonas aeruginosa Re(I)(CO)3(alpha-diimine)(HisX)+ (X = 83, 107, 109, 124, 126)Cu(II) azurins.
Blanco-Rodriguez, A.M., Busby, M., Ronayne, K., Towrie, M., Gradinaru, C., Sudhamsu, J., Sykora, J., Hof, M., Zalis, S., Di Bilio, A.J., Crane, B.R., Gray, H.B., Vlcek, A.(2009) J Am Chem Soc 131: 11788-11800
- PubMed: 19639996
- DOI: https://doi.org/10.1021/ja902744s
- Primary Citation of Related Structures:
2I7S, 3IBO - PubMed Abstract:
Photoinduced relaxation processes of five structurally characterized Pseudomonas aeruginosa Re(I)(CO)(3)(alpha-diimine)(HisX) (X = 83, 107, 109, 124, 126)Cu(II) azurins have been investigated by time-resolved (ps-ns) IR spectroscopy and emission spectroscopy. Crystal structures reveal the presence of Re-azurin dimers and trimers that in two cases (X = 107, 124) involve van der Waals interactions between interdigitated diimine aromatic rings. Time-dependent emission anisotropy measurements confirm that the proteins aggregate in mM solutions (D(2)O, KP(i) buffer, pD = 7.1). Excited-state DFT calculations show that extensive charge redistribution in the Re(I)(CO)(3) --> diimine (3)MLCT state occurs: excitation of this (3)MLCT state triggers several relaxation processes in Re-azurins whose kinetics strongly depend on the location of the metallolabel on the protein surface. Relaxation is manifested by dynamic blue shifts of excited-state nu(CO) IR bands that occur with triexponential kinetics: intramolecular vibrational redistribution together with vibrational and solvent relaxation give rise to subps, approximately 2, and 8-20 ps components, while the approximately 10(2) ps kinetics are attributed to displacement (reorientation) of the Re(I)(CO)(3)(phen)(im) unit relative to the peptide chain, which optimizes Coulombic interactions of the Re(I) excited-state electron density with solvated peptide groups. Evidence also suggests that additional segmental movements of Re-bearing beta-strands occur without perturbing the reaction field or interactions with the peptide. Our work demonstrates that time-resolved IR spectroscopy and emission anisotropy of Re(I) carbonyl-diimine complexes are powerful probes of molecular dynamics at or around the surfaces of proteins and protein-protein interfacial regions.
Organizational Affiliation:
School of Biological and Chemical Sciences, Queen Mary, University of London, Mile End Road, London E1 4NS, United Kingdom.