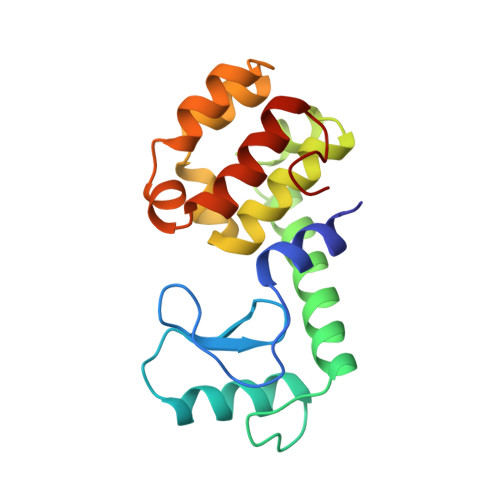
Energetic cost and structural consequences of burying a hydroxyl group within the core of a protein determined from Ala-->Ser and Val-->Thr substitutions in T4 lysozyme.
Blaber, M., Lindstrom, J.D., Gassner, N., Xu, J., Heinz, D.W., Matthews, B.W.(1993) Biochemistry 32: 11363-11373
- PubMed: 8218201
- DOI: https://doi.org/10.1021/bi00093a013
- Primary Citation of Related Structures:
118L, 119L, 120L, 122L, 123L, 125L, 126L, 127L, 128L, 206L, 221L, 224L - PubMed Abstract:
In order to determine the thermodynamic cost of introducing a polar group within the core of a protein, a series of nine Ala-->Ser and 3 Val-->Thr substitutions was constructed in T4 lysozyme. The sites were all within alpha-helices but ranged from fully solvent-exposed to totally buried. The range of destabilization incurred by the Ala-->Ser substitutions was found to be very similar to that for the Val-->Thr replacements. For the solvent-exposed and partly exposed sites the destabilization was modest (approximately less than 0.5 kcal/mol). For the completely buried sites the destabilization was larger, but variable (approximately 1-3 kcal/mol). Crystal structure determinations showed that the Ala-->Ser mutant structures were, in general, very similar to their wild-type counterparts, even though the replacements introduce a hydroxyl group. This is in part because the introduced serines are all within alpha-helices and at congested sites can avoid steric clashes with surrounding atoms by making a hydrogen bond to a backbone carbonyl oxygen in the preceding turn of the helix. The three substituted threonine side chains essentially superimpose on their valine counterparts but display somewhat larger conformational adjustments. The results illustrate how a protein structure will adapt in different ways to avoid the presence of an unsatisfied hydrogen bond donor or acceptor. In the most extreme case, Val 149-->Thr, which is also the most destabilizing variant (delta delta G = 2.8 kcal/mol), a water molecule is incorporated in the mutant structure in order to provide a hydrogen-bonding partner. The results are consistent with the view that many hydrogen bonds within proteins contribute only marginally to stability but that noncharged polar groups that lack a hydrogen-bonding partner are very destabilizing (delta delta G approximately greater than 3 kcal/mol). Supportive of other studies, the alpha-helix propensity of alanine is seen to be higher than that of serine (delta delta G = 0.46 +/- 0.04 kcal/mol), while threonine and valine are similar in alpha-helix propensity.
Organizational Affiliation:
Institute of Molecular Biology, Howard Hughes Medical Institute, Eugene, Oregon.