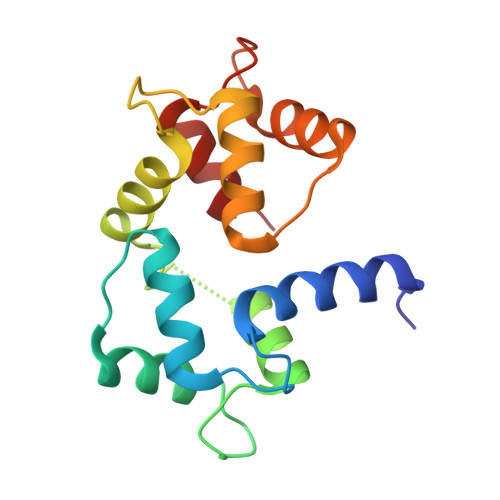
The structure of the complex of calmodulin with KAR-2: a novel mode of binding explains the unique pharmacology of the drug
Horvath, I., Harmat, V., Perczel, A., Palfi, V., Nyitrai, L., Nagy, A., Hlavanda, E., Naray-Szabo, G., Ovadi, J.(2005) J Biol Chem 280: 8266-8274
- PubMed: 15596444
- DOI: https://doi.org/10.1074/jbc.M410353200
- Primary Citation of Related Structures:
1XA5 - PubMed Abstract:
3'-(beta-Chloroethyl)-2',4'-dioxo-3,5'-spiro-oxazolidino-4-deacetoxyvinblastine (KAR-2) is a potent anti-microtubular agent that arrests mitosis in cancer cells without significant toxic side effects. In this study we demonstrate that in addition to targeting microtubules, KAR-2 also binds calmodulin, thereby countering the antagonistic effects of trifluoperazine. To determine the basis of both properties of KAR-2, the three-dimensional structure of its complex with Ca(2+)-calmodulin has been characterized both in solution using NMR and when crystallized using x-ray diffraction. Heterocorrelation ((1)H-(15)N heteronuclear single quantum coherence) spectra of (15)N-labeled calmodulin indicate a global conformation change (closure) of the protein upon its binding to KAR-2. The crystal structure at 2.12-A resolution reveals a more complete picture; KAR-2 binds to a novel structure created by amino acid residues of both the N- and C-terminal domains of calmodulin. Although first detected by x-ray diffraction of the crystallized ternary complex, this conformational change is consistent with its solution structure as characterized by NMR spectroscopy. It is noteworthy that a similar tertiary complex forms when calmodulin binds KAR-2 as when it binds trifluoperazine, even though the two ligands contact (for the most part) different amino acid residues. These observations explain the specificity of KAR-2 as an anti-microtubular agent; the drug interacts with a novel drug binding domain on calmodulin. Consequently, KAR-2 does not prevent calmodulin from binding most of its physiological targets.
Organizational Affiliation:
Institute of Enzymology, Biological Research Center, Hungarian Academy of Sciences, Karolina út 29 Budapest, H-1113 Hungary.