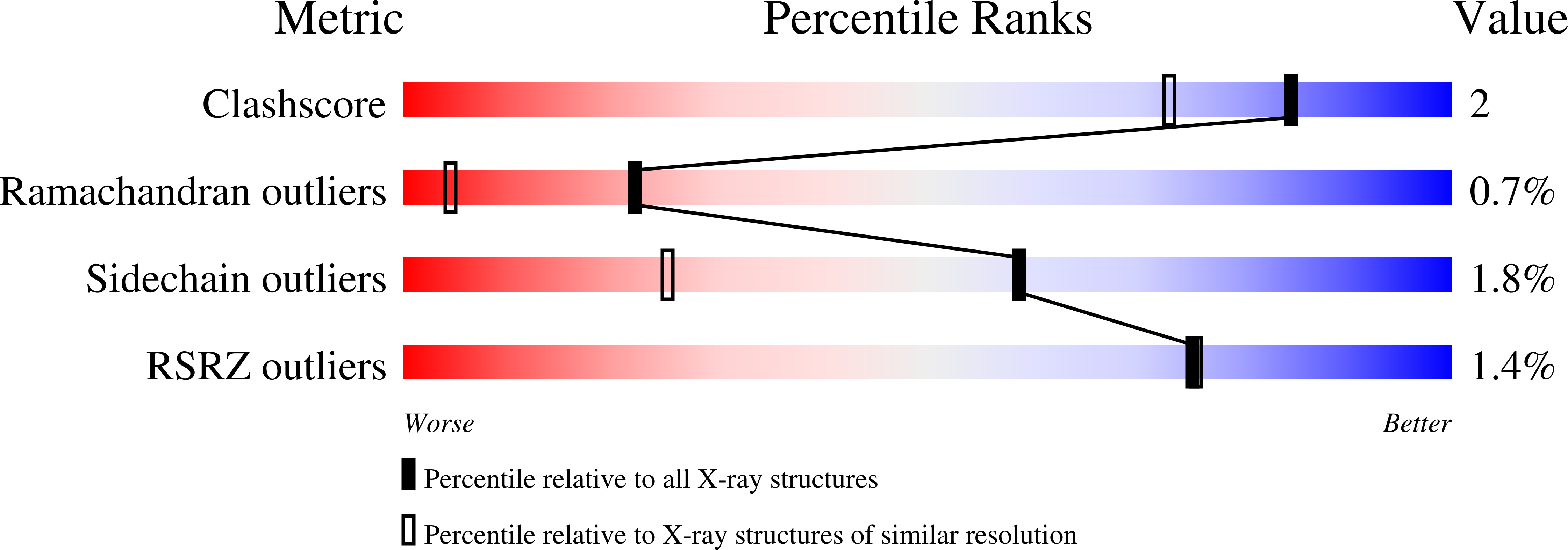
Comparison of the refined crystal structures of wild-type (1.34 A) flavodoxin from Desulfovibrio vulgaris and the S35C mutant (1.44 A) at 100 K.
Artali, R., Bombieri, G., Meneghetti, F., Gilardi, G., Sadeghi, S.J., Cavazzini, D., Rossi, G.L.(2002) Acta Crystallogr D Biol Crystallogr 58: 1787-1792
- PubMed: 12351822
- DOI: https://doi.org/10.1107/s0907444902012234
- Primary Citation of Related Structures:
1J8Q, 1J9E, 1J9G - PubMed Abstract:
Engineered flavodoxins in which a surface residue has been replaced by an exposed cysteine are useful modules to link multi-domain redox proteins obtained by gene fusion to electrode surfaces. In the present work, the crystal structure of the S35C mutant of Desulfovibrio vulgaris flavodoxin in the oxidized state has been determined and compared with a refined structure of the wild type (wt). The structure of wt flavodoxin (space group P4(3)2(1)2, unit-cell parameters a = 50.52, b = 50.52, c = 138.59 A) at 1.34 A resolution has been refined to R = 0.16 and R(free) = 0.18. The structure of the S35C mutant (space group P4(3)2(1)2, unit-cell parameters a = 50.55, b = 50.55, c = 138.39 A) at 1.44 A resolution has been refined to R = 0.13 and R(free) = 0.16. Data sets were collected with synchrotron radiation at 100 K. In the S35C mutant, the Cys35 thiol group points towards a hydrophobic region, whilst in the wt the Ser35 hydroxyl group points towards a more polar region. The solvent exposure of Cys35 is 43 A(2), of which 8 A(2) is for the sulfur. This is comparable to the exposure of 48 A(2) found for the wt Ser35, where that of the hydroxyl oxygen is also 8 A(2).
Organizational Affiliation:
Institute of Pharmaceutical Chemistry, University of Milano, Italy.