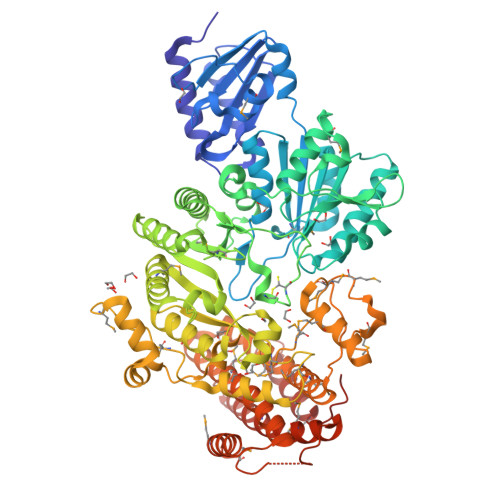
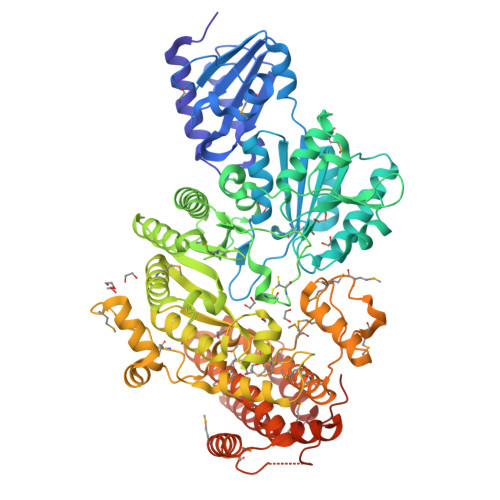
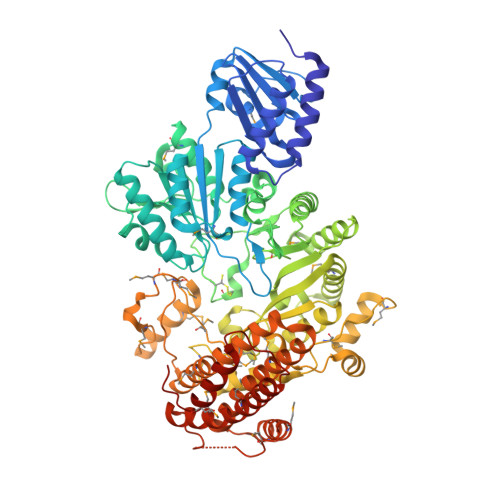
A B12-dependent radical SAM enzyme involved in oxetanocin A biosynthesis.
Bridwell-Rabb, J., Zhong, A., Sun, H.G., Drennan, C.L., Liu, H.W.(2017) Nature 544: 322-326
- PubMed: 28346939
- DOI: https://doi.org/10.1038/nature21689
- Primary Citation of Related Structures:
5UL2, 5UL3, 5UL4 - PubMed Abstract:
Oxetanocin A (OXT-A) is a potent antitumour, antiviral and antibacterial compound. Biosynthesis of OXT-A has been linked to a plasmid-borne Bacillus megaterium gene cluster that contains four genes: oxsA, oxsB, oxrA and oxrB. Here we show that both the oxsA and oxsB genes are required for the production of OXT-A. Biochemical analysis of the encoded proteins, a cobalamin (Cbl)-dependent S-adenosylmethionine (AdoMet) radical enzyme, OxsB, and an HD-domain phosphohydrolase, OxsA, reveals that OXT-A is derived from a 2'-deoxyadenosine phosphate in an OxsB-catalysed ring contraction reaction initiated by hydrogen atom abstraction from C2'. Hence, OxsB represents the first biochemically characterized non-methylating Cbl-dependent AdoMet radical enzyme. X-ray analysis of OxsB reveals the fold of a Cbl-dependent AdoMet radical enzyme, a family of enzymes with an estimated 7,000 members. Overall, this work provides a framework for understanding the interplay of AdoMet and Cbl cofactors and expands the catalytic repertoire of Cbl-dependent AdoMet radical enzymes.
Organizational Affiliation:
Howard Hughes Medical Institute, Massachusetts Institute of Technology, 77 Massachusetts Avenue, Cambridge, Massachusetts 02139, USA.