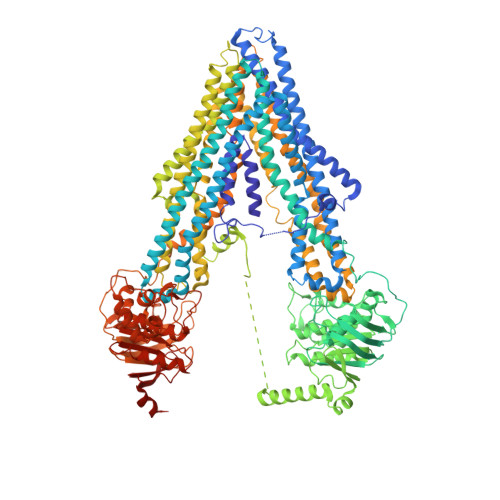
Crystal structure of the multidrug transporter P-glycoprotein from Caenorhabditis elegans.
Jin, M.S., Oldham, M.L., Zhang, Q., Chen, J.(2012) Nature 490: 566-569
- PubMed: 23000902
- DOI: https://doi.org/10.1038/nature11448
- Primary Citation of Related Structures:
4F4C - PubMed Abstract:
P-glycoprotein (P-gp) is an ATP-binding cassette transporter that confers multidrug resistance in cancer cells. It also affects the absorption, distribution and clearance of cancer-unrelated drugs and xenobiotics. For these reasons, the structure and function of P-gp have been studied extensively for decades. Here we present biochemical characterization of P-gp from Caenorhabditis elegans and its crystal structure at a resolution of 3.4 ångströms. We find that the apparent affinities of P-gp for anticancer drugs actinomycin D and paclitaxel are approximately 4,000 and 100 times higher, respectively, in the membrane bilayer than in detergent. This affinity enhancement highlights the importance of membrane partitioning when a drug accesses the transporter in the membrane. Furthermore, the transporter in the crystal structure opens its drug pathway at the level of the membrane's inner leaflet. In the helices flanking the opening to the membrane, we observe extended loops that may mediate drug binding, function as hinges to gate the pathway or both. We also find that the interface between the transmembrane and nucleotide-binding domains, which couples ATP hydrolysis to transport, contains a ball-and-socket joint and salt bridges similar to the ATP-binding cassette importers, suggesting that ATP-binding cassette exporters and importers may use similar mechanisms to achieve alternating access for transport. Finally, a model of human P-gp derived from the structure of C. elegans P-gp not only is compatible with decades of biochemical analysis, but also helps to explain perplexing functional data regarding the Phe335Ala mutant. These results increase our understanding of the structure and function of this important molecule.
- Department of Biological Sciences, Purdue University, Indiana 47907, USA.
Organizational Affiliation: