Structural Insights into the Mechanism of the PLP Synthase Holoenzyme from Thermotoga maritima
Zein, F., Zhang, Y., Kang, Y.N., Burns, K., Begley, T.P., Ealick, S.E.(2006) Biochemistry 45: 14609-14620
- PubMed: 17144654
- DOI: https://doi.org/10.1021/bi061464y
- Primary Citation of Related Structures:
2ISS - PubMed Abstract:
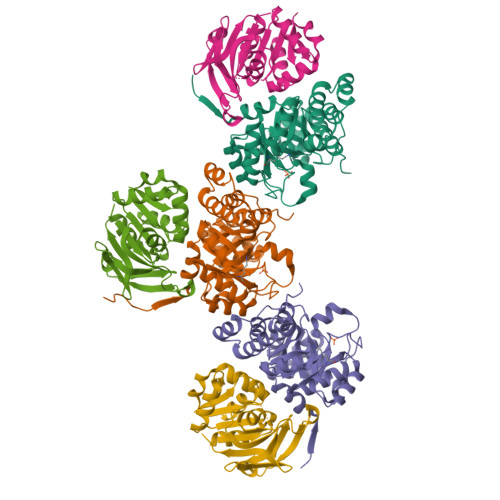
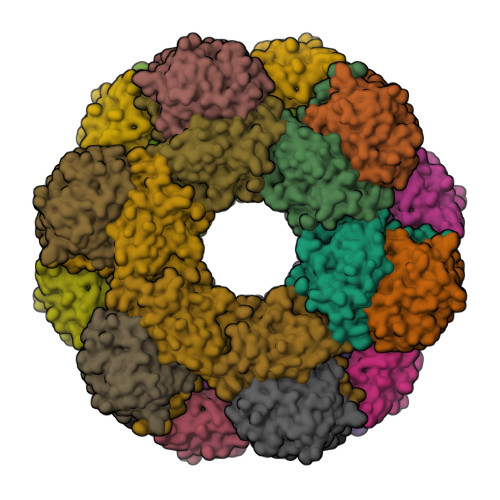
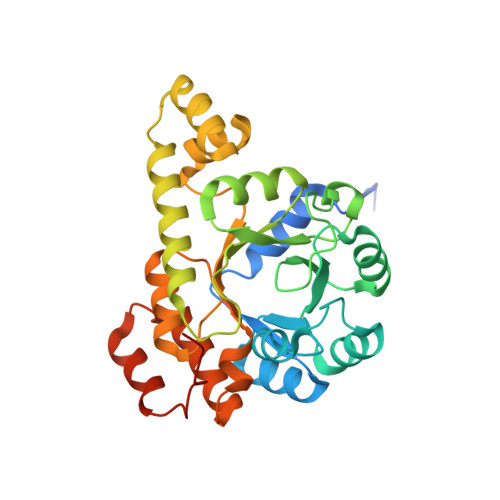
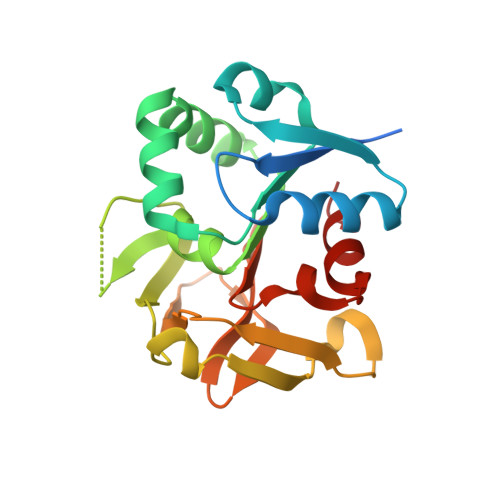
Pyridoxal 5'-phosphate (PLP) is the biologically active form of vitamin B6 and is an important cofactor for several of the enzymes involved in the metabolism of amine-containing natural products such as amino acids and amino sugars. The PLP synthase holoenzyme consists of two subunits: YaaD catalyzes the condensation of ribulose 5-phosphate, glyceraldehyde-3-phosphate, and ammonia, and YaaE catalyzes the production of ammonia from glutamine. Here we describe the structure of the PLP synthase complex (YaaD-YaaE) from Thermotoga maritima at 2.9 A resolution. This complex consists of a core of 12 YaaD monomers with 12 noninteracting YaaE monomers attached to the core. Compared with the previously published structure of PdxS (a YaaD ortholog in Geobacillus stearothermophilus), the N-terminus (1-18), which includes helix alpha0, the beta2-alpha2 loop (46-56), which includes new helix alpha2a, and the C-terminus (270-280) of YaaD are ordered in the complex but disordered in PdxS. A ribulose 5-phosphate is bound to YaaD via an imine with Lys82. Previous studies have demonstrated a similar imine at Lys149 and not at Lys81 (equivalent to Lys150 and Lys82 in T. maritima) for the Bacillus subtilis enzyme suggesting the possibility that two separate sites on YaaD are involved in PLP formation. A phosphate from the crystallization solution is found bound to YaaD and also serves as a marker for a possible second active site. An ammonia channel that connects the active site of YaaE with the ribulose 5-phosphate binding site was identified. This channel is similar to one found in imidazole glycerol phosphate synthase; however, when the beta-barrels of the two complexes are superimposed, the glutaminase domains are rotated by about 180 degrees with respect to each other.
Organizational Affiliation:
Department of Chemistry and Chemical Biology, Cornell University, Ithaca, New York 14853, USA.