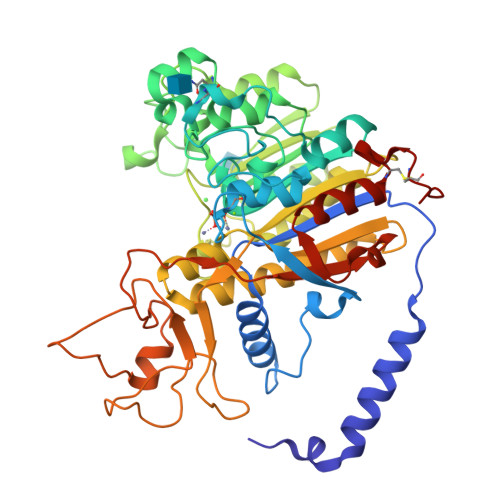
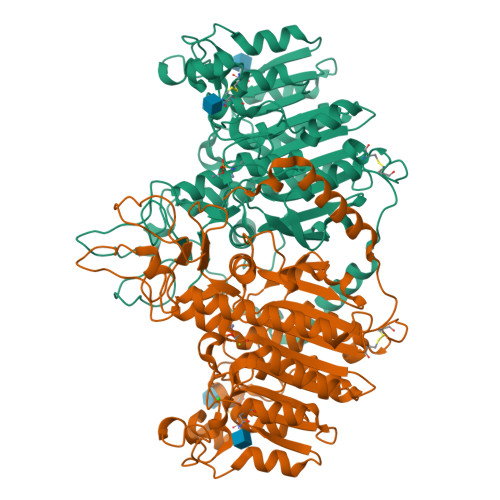
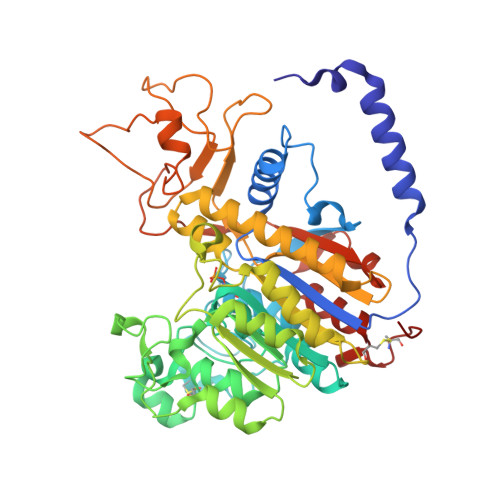
Structural studies of human alkaline phosphatase in complex with strontium: Implication for its secondary effect in bones.
Llinas, P., Masella, M., Stigbrand, T., Menez, A., Stura, E.A., Le Du, M.H.(2006) Protein Sci 15: 1691-1700
- PubMed: 16815919
- DOI: https://doi.org/10.1110/ps.062123806
- Primary Citation of Related Structures:
2GLQ - PubMed Abstract:
Strontium is used in the treatment of osteoporosis as a ranelate compound, and in the treatment of painful scattered bone metastases as isotope. At very high doses and in certain conditions, it can lead to osteomalacia characterized by impairment of bone mineralization. The osteomalacia symptoms resemble those of hypophosphatasia, a rare inherited disorder associated with mutations in the gene encoding for tissue-nonspecific alkaline phosphatase (TNAP). Human alkaline phosphatases have four metal binding sites--two for zinc, one for magnesium, and one for calcium ion--that can be substituted by strontium. Here we present the crystal structure of strontium-substituted human placental alkaline phosphatase (PLAP), a related isozyme of TNAP, in which such replacement can have important physiological implications. The structure shows that strontium substitutes the calcium ion with concomitant modification of the metal coordination. The use of the flexible and polarizable force-field TCPEp (topological and classical polarization effects for proteins) predicts that calcium or strontium has similar interaction energies at the calcium-binding site of PLAP. Since calcium helps stabilize a large area that includes loops 210-228 and 250-297, its substitution by strontium could affect the stability of this region. Energy calculations suggest that only at high doses of strontium, comparable to those found for calcium, can strontium substitute for calcium. Since osteomalacia is observed after ingestion of high doses of strontium, alkaline phosphatase is likely to be one of the targets of strontium, and thus this enzyme might be involved in this disease.
Organizational Affiliation:
Laboratoire de Structure des Protéines, Département d'Ingénierie et d'Etude des Protéines, Commissariat à l'Energie Atomique, Gif sur Yvette, France.