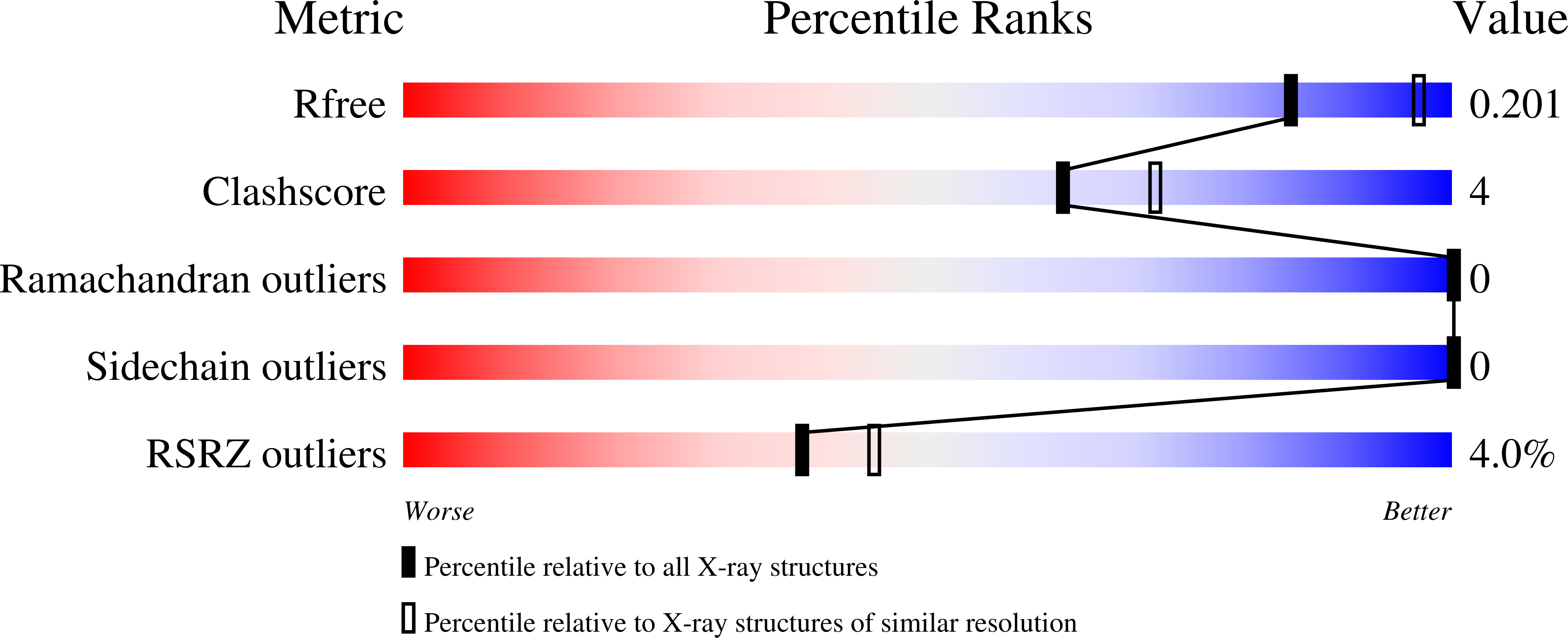
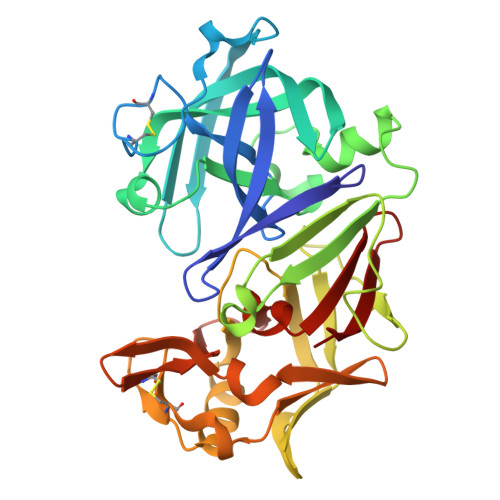
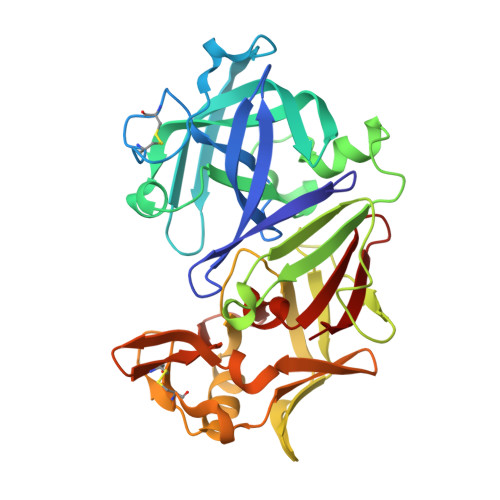
Effect of pH on the structure of rhizopuspepsin.
Prasad, B.V., Suguna, K.(2003) Acta Crystallogr D Biol Crystallogr 59: 1755-1761
- PubMed: 14501114
- DOI: https://doi.org/10.1107/s0907444903016068
- Primary Citation of Related Structures:
1UH7, 1UH8, 1UH9 - PubMed Abstract:
The crystal structure of rhizopuspepsin has been determined at three different pH values (4.6, 7.0 and 8.0) and compared with the previously reported structure at pH 6.0. A pH-sensitive region in the protein has been identified where certain structural changes take place at pH 8.0. An increase in the mobility of loops, weakening of hydrogen bonding and ionic interactions and a change in the water structure have been observed in this region. The loop between the first and the second beta-strands of the N-terminus shows increased mobility at high pH. This loop is known to be highly flexible in aspartic proteinases, aiding in relocating the N-terminal beta-strand segment in pH-related structural transformations. The observed changes in rhizopuspepsin indicate the triggering of a possible denatured state by high pH. The conformation of the active aspartates and the geometry of the catalytic site exhibit remarkable rigidity in this pH range.
Organizational Affiliation:
Molecular Biophysics Unit, Indian Institute of Science, Bangalore 560 012, India.