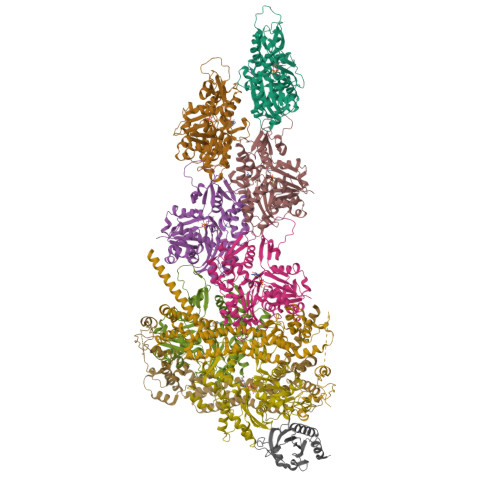
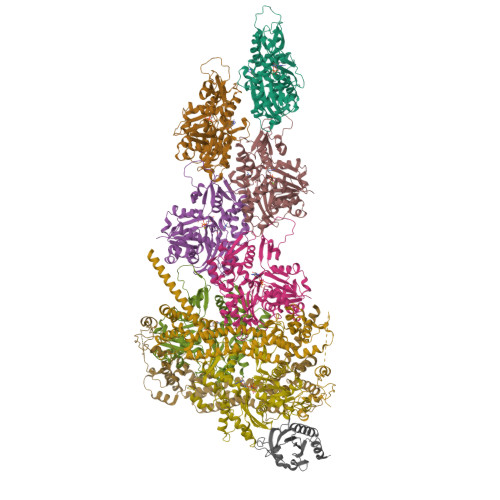
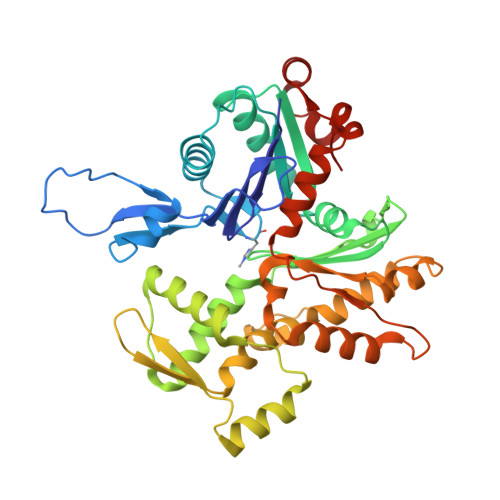
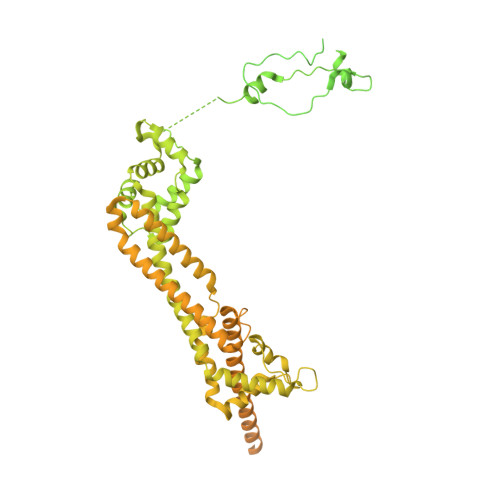
Mechanisms of actin filament severing and elongation by formins.
Palmer, N.J., Barrie, K.R., Dominguez, R.(2024) Nature 632: 437-442
- PubMed: 38843827
- DOI: https://doi.org/10.1038/s41586-024-07637-0
- Primary Citation of Related Structures:
9AZ4, 9AZP, 9AZQ, 9B03, 9B0K, 9B27, 9B3D - PubMed Abstract:
Humans express fifteen formins, playing crucial roles in actin-based processes, such as cytokinesis, cell motility, and mechanotransduction 1,2 . However, the lack of structures bound to the actin filament (F-actin) has been a major impediment to understanding formin function. While formins are known for their ability to nucleate and elongate F-actin 3-7 , some formins can additionally depolymerize, sever, or bundle F-actin. Two mammalian formins, inverted formin-2 (INF2) and diaphanous-1 (Dia1), exemplify this diversity. INF2 displays potent severing activity but elongates weakly 8-11 , whereas Dia1 has potent elongation activity but does not sever 4,8 . Using cryo-electron microscopy (cryo-EM), we reveal five structural states of INF2 and two of Dia1 bound to the middle and barbed end of F-actin. INF2 and Dia1 bind differently to these sites, consistent with their distinct activities. The FH2 and WH2 domains of INF2 are positioned to sever F-actin, whereas Dia1 appears unsuited for severing. Structures also show how profilin-actin is delivered to the fast-growing barbed end, and how this is followed by a transition of the incoming monomer into the F-actin conformation and the release of profilin. Combined, the seven structures presented here provide step-by-step visualization of the mechanisms of F-actin severing and elongation by formins.
Organizational Affiliation:
Department of Physiology and Biochemistry and Molecular Biophysics Graduate Group, University of Pennsylvania Perelman School of Medicine, Philadelphia, Pennsylvania, USA.