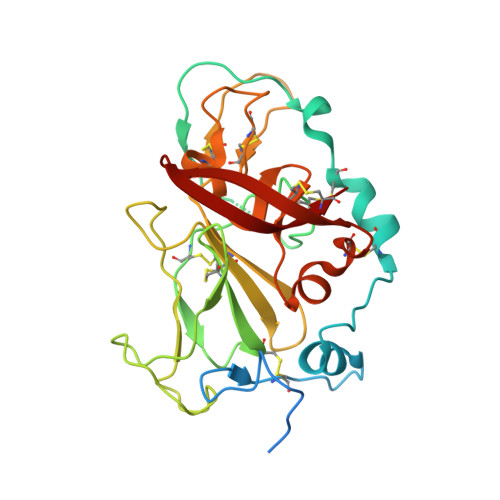
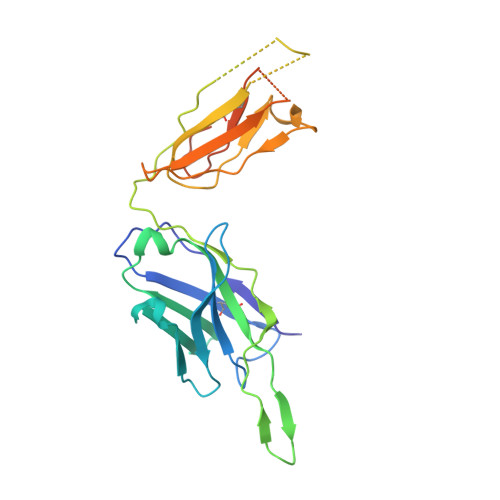
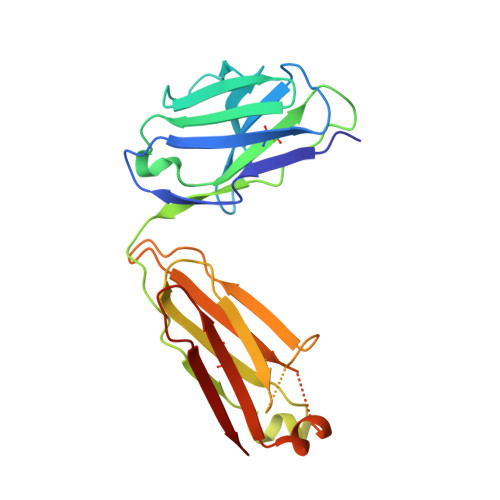
Convergent evolution and targeting of diverse E2 epitopes by human broadly neutralizing antibodies are associated with HCV clearance.
Ogega, C.O., Skinner, N.E., Schoenle, M.V., Wilcox, X.E., Frumento, N., Wright, D.A., Paul, H.T., Sinnis-Bourozikas, A., Clark, K.E., Figueroa, A., Bjorkman, P.J., Ray, S.C., Flyak, A.I., Bailey, J.R.(2024) Immunity 57: 890-903.e6
- PubMed: 38518779
- DOI: https://doi.org/10.1016/j.immuni.2024.03.001
- Primary Citation of Related Structures:
8W0V, 8W0W, 8W0X, 8W0Y - PubMed Abstract:
The early appearance of broadly neutralizing antibodies (bNAbs) in serum is associated with spontaneous hepatitis C virus (HCV) clearance, but to date, the majority of bNAbs have been isolated from chronically infected donors. Most of these bNAbs use the V H 1-69 gene segment and target the envelope glycoprotein E2 front layer. Here, we performed longitudinal B cell receptor (BCR) repertoire analysis on an elite neutralizer who spontaneously cleared multiple HCV infections. We isolated 10,680 E2-reactive B cells, performed BCR sequencing, characterized monoclonal B cell cultures, and isolated bNAbs. In contrast to what has been seen in chronically infected donors, the bNAbs used a variety of V H genes and targeted at least three distinct E2 antigenic sites, including sites previously thought to be non-neutralizing. Diverse front-layer-reactive bNAb lineages evolved convergently, acquiring breadth-enhancing somatic mutations. These findings demonstrate that HCV clearance-associated bNAbs are genetically diverse and bind distinct antigenic sites that should be the target of vaccine-induced bNAbs.
Organizational Affiliation:
Division of Infectious Diseases, Department of Medicine, Johns Hopkins University School of Medicine, Baltimore, MD, USA; Department of Pharmacology and Molecular Sciences, Johns Hopkins University School of Medicine, Baltimore, MD, USA.