Structural and biochemical analysis of the PARP1-homology region of PARP4/vault PARP.
Frigon, L., Pascal, J.M.(2023) Nucleic Acids Res 51: 12492-12507
- PubMed: 37971310
- DOI: https://doi.org/10.1093/nar/gkad1064
- Primary Citation of Related Structures:
8SWY, 8SWZ, 8SX1, 8SX2 - PubMed Abstract:
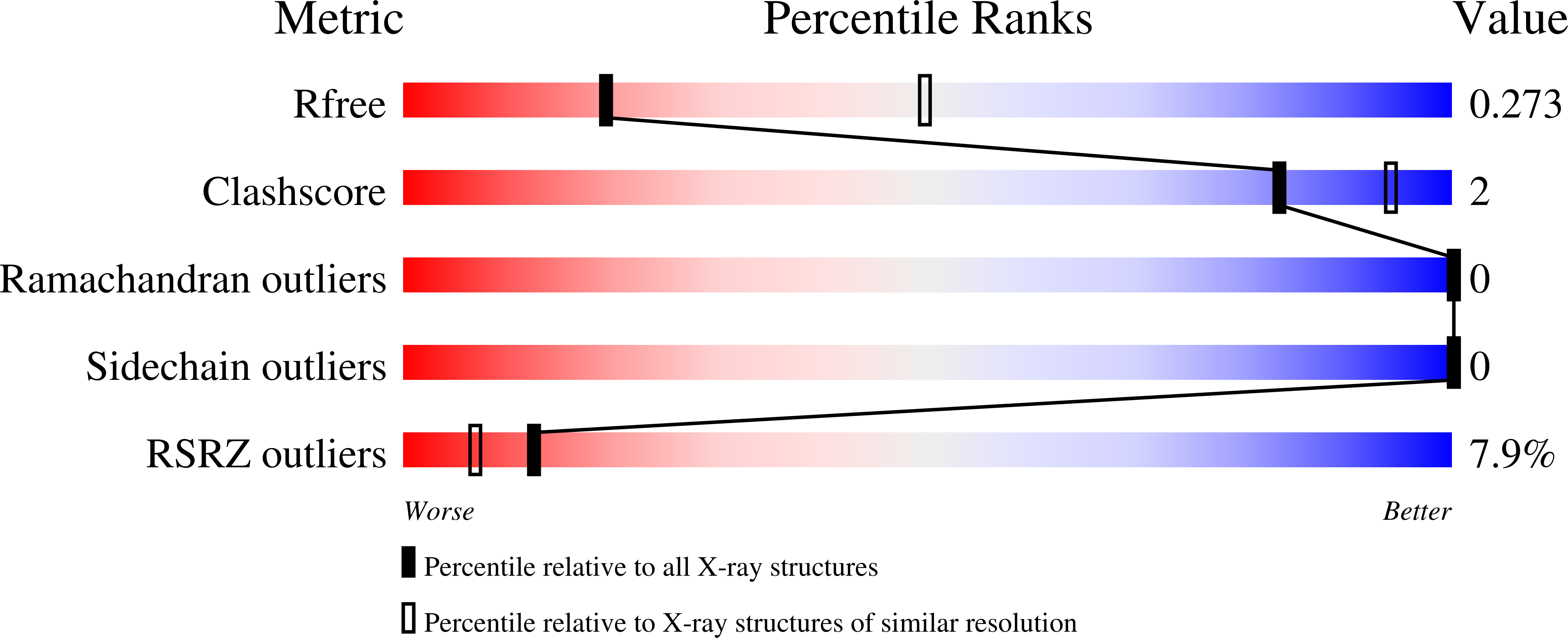
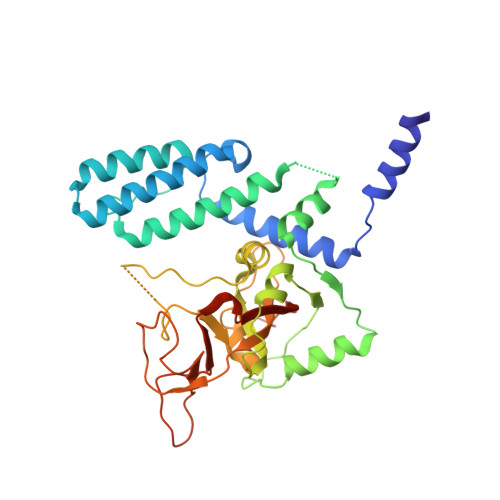
PARP4 is an ADP-ribosyltransferase that resides within the vault ribonucleoprotein organelle. Our knowledge of PARP4 structure and biochemistry is limited relative to other PARPs. PARP4 shares a region of homology with PARP1, an ADP-ribosyltransferase that produces poly(ADP-ribose) from NAD+ in response to binding DNA breaks. The PARP1-homology region of PARP4 includes a BRCT fold, a WGR domain, and the catalytic (CAT) domain. Here, we have determined X-ray structures of the PARP4 catalytic domain and performed biochemical analysis that together indicate an active site that is open to NAD+ interaction, in contrast to the closed conformation of the PARP1 catalytic domain that blocks access to substrate NAD+. We have also determined crystal structures of the minimal ADP-ribosyltransferase fold of PARP4 that illustrate active site alterations that restrict PARP4 to mono(ADP-ribose) rather than poly(ADP-ribose) modifications. We demonstrate that PARP4 interacts with vault RNA, and that the BRCT is primarily responsible for the interaction. However, the interaction does not lead to stimulation of mono(ADP-ribosylation) activity. The BRCT-WGR-CAT of PARP4 has lower activity than the CAT alone, suggesting that the BRCT and WGR domains regulate catalytic output. Our study provides first insights into PARP4 structure and regulation and expands understanding of PARP structural biochemistry.
Organizational Affiliation:
Department of Biochemistry and Molecular Medicine, Université de Montréal, Montréal, Qc H3T 1J4, Canada.