Kinetic and structural details of urease inactivation by thiuram disulphides.
Mazzei, L., Paul, A., Cianci, M., Devodier, M., Mandelli, D., Carloni, P., Ciurli, S.(2023) J Inorg Biochem 250: 112398-112398
- PubMed: 37879152
- DOI: https://doi.org/10.1016/j.jinorgbio.2023.112398
- Primary Citation of Related Structures:
8Q2E - PubMed Abstract:
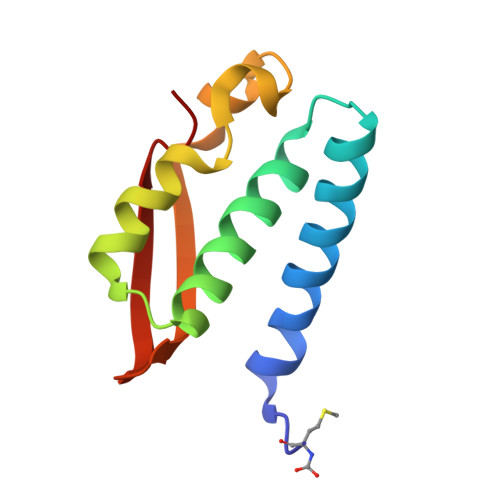
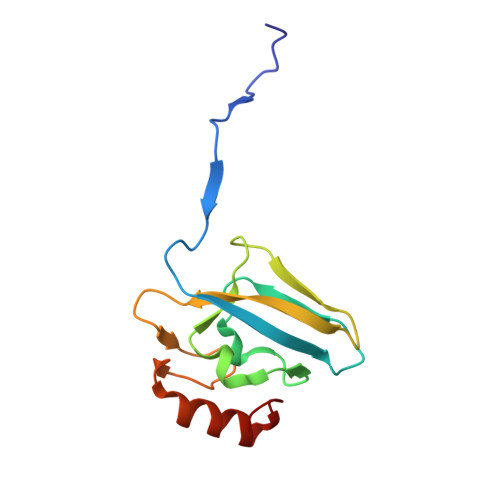
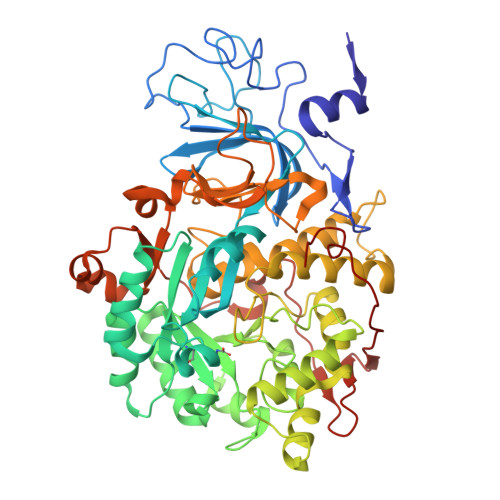
This paper reports on the molecular details of the reactivity of urease, a nickel-dependent enzyme that catalyses the last step of organic nitrogen mineralization, with thiuram disulphides, a class of molecules known to inactivate the enzyme with high efficacy but for which the mechanism of action had not been yet established. IC 50 values of tetramethylthiuram disulphide (TMTD or Thiram) and tetraethylthiuram disulphide (TETD or Disulfiram) in the low micromolar range were determined for plant and bacterial ureases. The X-ray crystal structure of Sporosarcina pasteurii urease inactivated by Thiram, determined at 1.68 Å resolution, revealed the presence of a covalent modification of the catalytically essential cysteine residue. This is located on the flexible flap that modulates the size of the active site channel and cavity. Formation of a Cys-S-S-C(S)-N(CH 3 ) 2 functionality responsible for enzyme inactivation was observed. Quantum-mechanical calculations carried out to rationalise the large reactivity of the active site cysteine support the view that a conserved histidine residue, adjacent to the cysteine in the active site flap, modulates the charge and electron density along the thiol SH bond by shifting electrons towards the sulphur atom and rendering the thiol proton more reactive. We speculate that this proton could be transferred to the nickel-coordinated urea amide group to yield a molecule of ammonia from the generated C urea -NH 3 + functionality during catalysis.
Organizational Affiliation:
Laboratory of Bioinorganic Chemistry, Department of Pharmacy and Biotechnology (FaBiT), University of Bologna, Viale Giuseppe Fanin 40, Bologna I-40127, Italy. Electronic address: luca.mazzei2@unibo.it.