A novel allosteric site employs a conserved inhibition mechanism in human kidney-type glutaminase.
Shankar, S., Ramachandran, S., Tulsian, N., Radhakrishnan, S., Jobichen, C., Sivaraman, J.(2023) FEBS J 290: 2437-2448
- PubMed: 36259273
- DOI: https://doi.org/10.1111/febs.16658
- Primary Citation of Related Structures:
8GWR - PubMed Abstract:
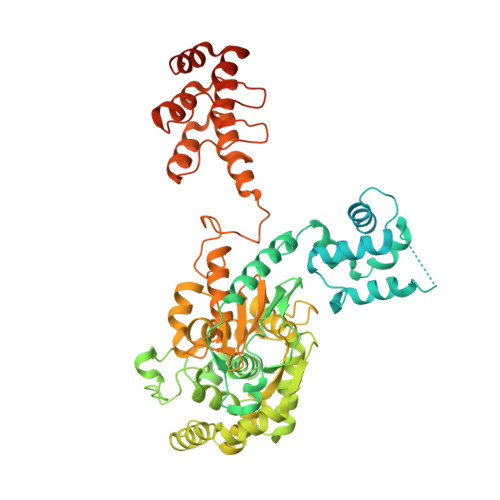
Glutaminase catalyses the metabolic process called glutaminolysis. Cancer cells harness glutaminolysis to increase energy reserves under stressful conditions for rapid proliferation. Glutaminases are upregulated in many tumours. In humans, the kidney-type glutaminase (KGA) isoform is highly expressed in the kidney, brain, intestine, foetal liver, lymphocytes and in many tumours. Glutaminase inhibition is shown to be effective in controlling cancers. Previously, we and others reported the inhibition mechanism of KGA using various inhibitors that target the active and allosteric sites of the enzyme. Here, we report the identification of a novel allosteric site in KGA using the compound DDP through its complex crystal structure combined with mutational and hydrogen-deuterium exchange mass spectrometry studies. This allosteric site is located at the dimer interface, situated ~ 31 Å away from the previously identified allosteric site and ~ 32 Å away from the active site. Remarkably, the mechanism of inhibition is conserved, irrespective of which allosteric pocket is targeted, causing the same conformational changes in the key loop near the active site (Glu312-Pro329) and subsequent enzyme inactivation. Contrary to the previously identified allosteric site, the identified new allosteric site is primarily hydrophilic. This site could be effectively targeted for the synthesis of specific and potent water-soluble inhibitors of glutaminase, which will lead to the development of anticancer drugs.
Organizational Affiliation:
Department of Biological Sciences, National University of Singapore, Singapore.