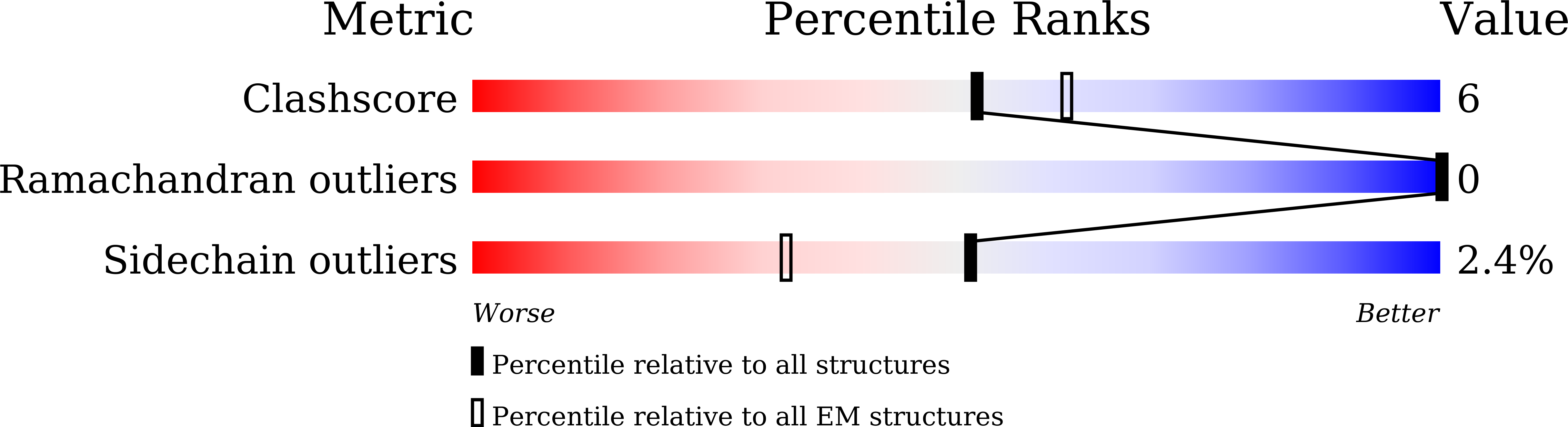Cryo-EM structure of GABA transporter 1 reveals substrate recognition and transport mechanism.
Nayak, S.R., Joseph, D., Hofner, G., Dakua, A., Athreya, A., Wanner, K.T., Kanner, B.I., Penmatsa, A.(2023) Nat Struct Mol Biol 30: 1023-1032
- PubMed: 37400654
- DOI: https://doi.org/10.1038/s41594-023-01011-w
- Primary Citation of Related Structures:
8GNK - PubMed Abstract:
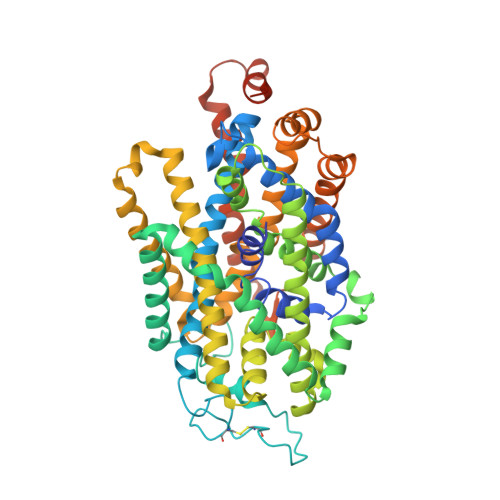
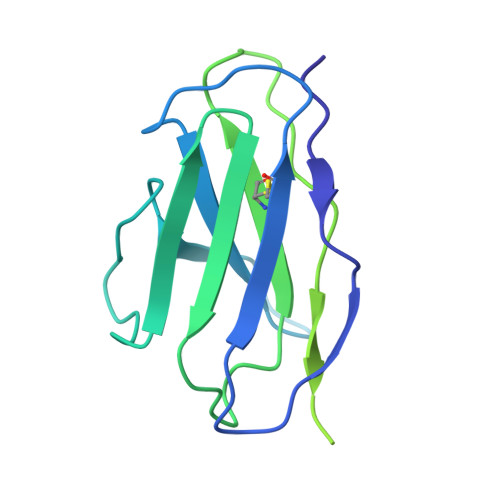
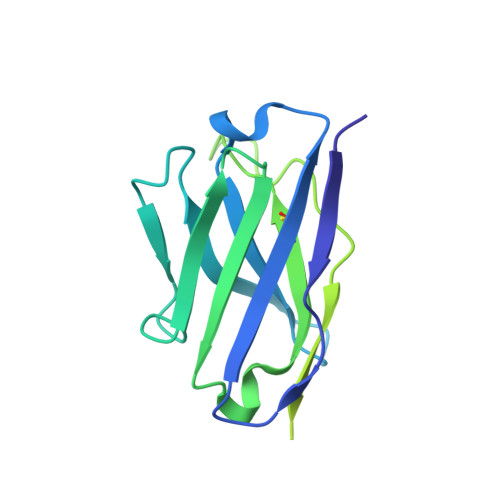
The inhibitory neurotransmitter γ-aminobutyric acid (GABA) is cleared from the synaptic cleft by the sodium- and chloride-coupled GABA transporter GAT1. Inhibition of GAT1 prolongs the GABAergic signaling at the synapse and is a strategy to treat certain forms of epilepsy. In this study, we present the cryo-electron microscopy structure of Rattus norvegicus GABA transporter 1 (rGAT1) at a resolution of 3.1 Å. The structure elucidation was facilitated by epitope transfer of a fragment-antigen binding (Fab) interaction site from the Drosophila dopamine transporter (dDAT) to rGAT1. The structure reveals rGAT1 in a cytosol-facing conformation, with a linear density in the primary binding site that accommodates a molecule of GABA, a displaced ion density proximal to Na site 1 and a bound chloride ion. A unique insertion in TM10 aids the formation of a compact, closed extracellular gate. Besides yielding mechanistic insights into ion and substrate recognition, our study will enable the rational design of specific antiepileptics.
Organizational Affiliation:
Molecular Biophysics Unit, Indian Institute of Science, Bangalore, India.