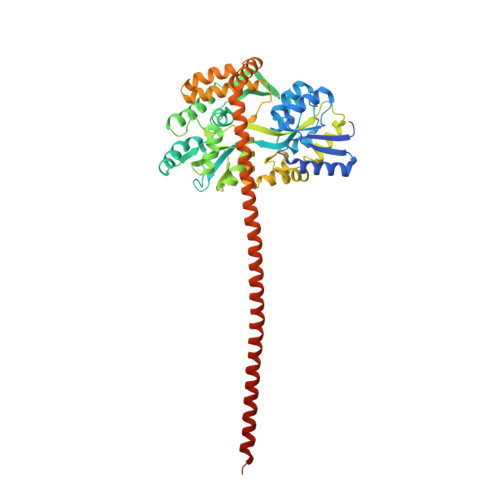
A novel single alpha-helix DNA-binding domain in CAF-1 promotes gene silencing and DNA damage survival through tetrasome-length DNA selectivity and spacer function.
Rosas, R., Aguilar, R.R., Arslanovic, N., Seck, A., Smith, D.J., Tyler, J.K., Churchill, M.E.A.(2023) Elife 12
- PubMed: 37432722
- DOI: https://doi.org/10.7554/eLife.83538
- Primary Citation of Related Structures:
8DEI - PubMed Abstract:
The histone chaperone chromatin assembly factor 1 (CAF-1) deposits two nascent histone H3/H4 dimers onto newly replicated DNA forming the central core of the nucleosome known as the tetrasome. How CAF-1 ensures there is sufficient space for the assembly of tetrasomes remains unknown. Structural and biophysical characterization of the lysine/glutamic acid/arginine-rich (KER) region of CAF-1 revealed a 128-Å single alpha-helix (SAH) motif with unprecedented DNA-binding properties. Distinct KER sequence features and length of the SAH drive the selectivity of CAF-1 for tetrasome-length DNA and facilitate function in budding yeast. In vivo, the KER cooperates with the DNA-binding winged helix domain in CAF-1 to overcome DNA damage sensitivity and maintain silencing of gene expression. We propose that the KER SAH links functional domains within CAF-1 with structural precision, acting as a DNA-binding spacer element during chromatin assembly.
Organizational Affiliation:
Program in Structural Biology and Biochemistry, University of Colorado Anschutz Medical Campus, Aurora, United States.