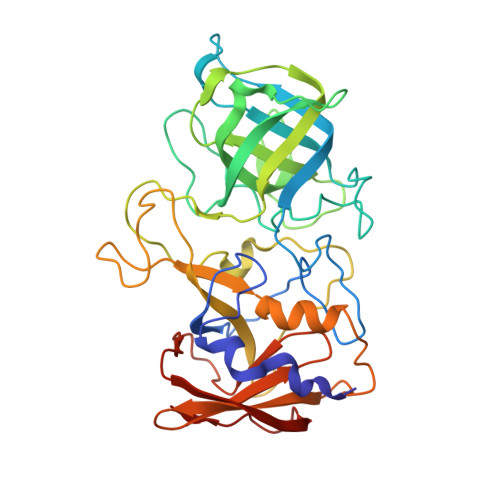
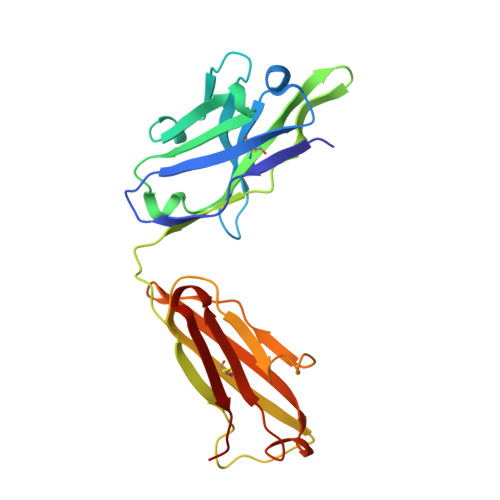
Structural Basis for Rabbit Hemorrhagic Disease Virus Antibody Specificity.
Leuthold, M.M., Kilic, T., Devant, J.M., Landeta, O., Parra, F., Dalton, K.P., Hansman, G.S.(2022) J Virol 96: e0121722-e0121722
- PubMed: 36326275
- DOI: https://doi.org/10.1128/jvi.01217-22
- Primary Citation of Related Structures:
8CYL, 8CZ5 - PubMed Abstract:
Rabbit hemorrhagic disease virus (RHDV) typically causes a fatal disease in rabbits. In Australia, RHDV was imported to control the feral rabbit population, while it poses a severe threat to native rabbits in other countries. RHDV variants are genetically diverse and serological studies using antibodies isolated from infected rabbits or raised against RHDV virus-like particles (VLPs) have found RHDV variants antigenically distinct. In this study, we determined the X-ray crystal structure of an RHDV GI.2 (N11 strain) protruding (P) domain in complex with a diagnostic monoclonal antibody (2D9) Fab. We showed that 2D9 interacted with conserved and variable residues on top of the P domain with nanomolar affinity. To better illustrate 2D9 specificity, we determined the X-ray crystal structure of an RHDV GI.1b (Ast89 strain) that was a 2D9 non-binder. Structural analysis indicated that amino acid substitutions on the GI.1b P domain likely restricted 2D9 binding. Interestingly, a model of the GI.2 P domain-Fab complex superimposed onto a cryo-EM structure of an RHDV VLP revealed that 2D9 Fab molecules clashed with neighboring Fabs and indicated that there was a reduced antibody binding occupancy. Moreover, the RHDV GI.2 histo-blood group antigen (HBGA) co-factor binding site appeared obstructed when 2D9 was modeled on the VLP and suggested that 2D9 might also function by blocking HBGA attachment. Overall, this new data provides the first structural basis of RHDV antibody specificity and explains how amino acid variation at the binding site likely restricts 2D9 cross-reactivity. IMPORTANCE Isolated RHDV antibodies have been used for decades to distinguish between antigenic variants, monitor temporal capsid evolution, and examine neutralizing capacities. In this study, we provided the structural basis for an RHDV GI.2 specific diagnostic antibody (2D9) binding and reveal that a small number of amino acid substitutions at the binding site could differentiate between RHDV GI.2 and GI.1b. This novel structural information provides a framework for understanding how RHDV displays a specific antigenic epitope and engages an antibody at the atomic level. Importantly, part of the 2D9 binding region was earlier reported to contain a neutralizing epitope and our structural modeling as well as recent human norovirus antibody-mediated neutralization studies, suggest that the 2D9 antibody has the potential to block HBGA attachment. These new findings should aid in characterizing antigenic variants and advance the development of novel monoclonal antibodies for diagnostics and therapeutics.
Organizational Affiliation:
Medicinal Chemistry, Institute of Pharmacy and Molecular Biotechnology, University of Heidelberg, Heidelberg, Germany.