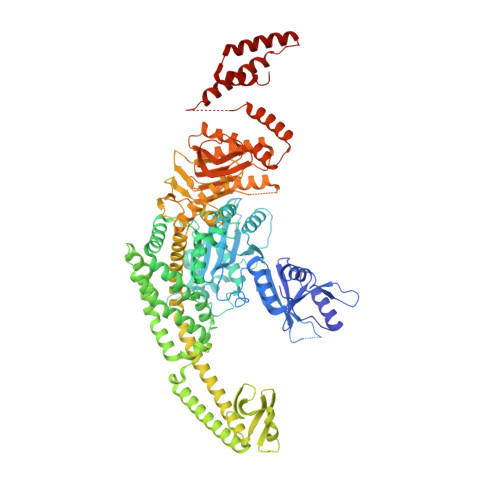
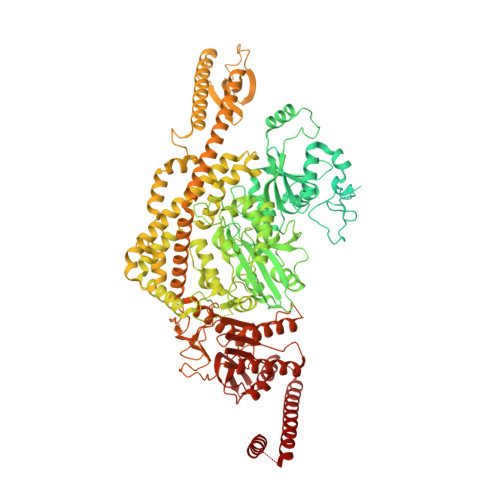
Unexpected moves: a conformational change in MutS alpha enables high-affinity DNA mismatch binding.
Bruekner, S.R., Pieters, W., Fish, A., Liaci, A.M., Scheffers, S., Rayner, E., Kaldenbach, D., Drost, L., Dekker, M., van Hees-Stuivenberg, S., Delzenne-Goette, E., de Konink, C., Houlleberghs, H., Dubbink, H.J., AlSaegh, A., de Wind, N., Forster, F., Te Riele, H., Sixma, T.K.(2023) Nucleic Acids Res 51: 1173-1188
- PubMed: 36715327
- DOI: https://doi.org/10.1093/nar/gkad015
- Primary Citation of Related Structures:
8AG6 - PubMed Abstract:
The DNA mismatch repair protein MutSα recognizes wrongly incorporated DNA bases and initiates their correction during DNA replication. Dysfunctions in mismatch repair lead to a predisposition to cancer. Here, we study the homozygous mutation V63E in MSH2 that was found in the germline of a patient with suspected constitutional mismatch repair deficiency syndrome who developed colorectal cancer before the age of 30. Characterization of the mutant in mouse models, as well as slippage and repair assays, shows a mildly pathogenic phenotype. Using cryogenic electron microscopy and surface plasmon resonance, we explored the mechanistic effect of this mutation on MutSα function. We discovered that V63E disrupts a previously unappreciated interface between the mismatch binding domains (MBDs) of MSH2 and MSH6 and leads to reduced DNA binding. Our research identifies this interface as a 'safety lock' that ensures high-affinity DNA binding to increase replication fidelity. Our mechanistic model explains the hypomorphic phenotype of the V63E patient mutation and other variants in the MBD interface.
Organizational Affiliation:
Division of Biochemistry, Netherlands Cancer Institute and Oncode Institute, 1066 CX Amsterdam, The Netherlands.