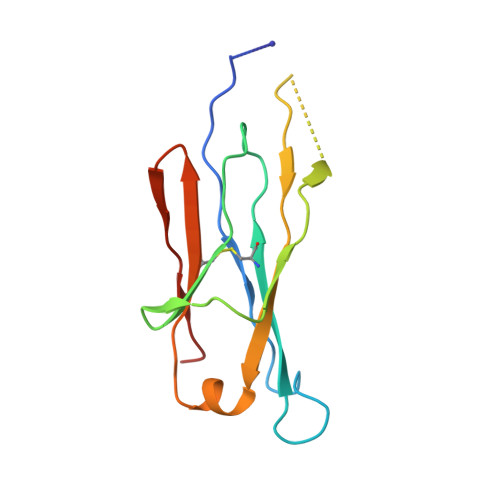
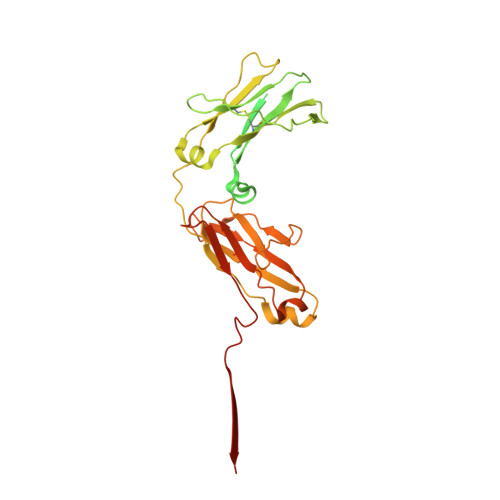
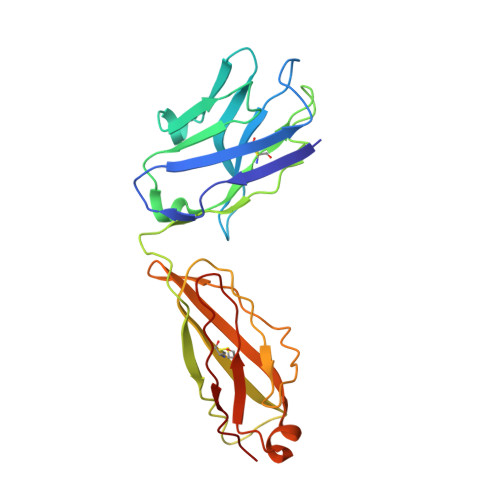
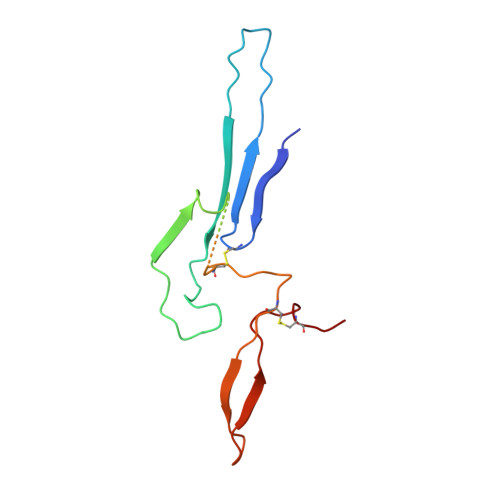
Cryomicroscopy reveals the structural basis for a flexible hinge motion in the immunoglobulin M pentamer.
Chen, Q., Menon, R., Calder, L.J., Tolar, P., Rosenthal, P.B.(2022) Nat Commun 13: 6314-6314
- PubMed: 36274064
- DOI: https://doi.org/10.1038/s41467-022-34090-2
- Primary Citation of Related Structures:
7QDO, 8ADY, 8ADZ, 8AE0, 8AE2, 8AE3 - PubMed Abstract:
Immunoglobulin M (IgM) is the most ancient of the five isotypes of immunoglobulin (Ig) molecules and serves as the first line of defence against pathogens. Here, we use cryo-EM to image the structure of the human full-length IgM pentamer, revealing antigen binding domains flexibly attached to the asymmetric and rigid core formed by the Cμ4 and Cμ3 constant regions and the J-chain. A hinge is located at the Cμ3/Cμ2 domain interface, allowing Fabs and Cμ2 to pivot as a unit both in-plane and out-of-plane. This motion is different from that observed in IgG and IgA, where the two Fab arms are able to swing independently. A biased orientation of one pair of Fab arms results from asymmetry in the constant domain (Cμ3) at the IgM subunit interacting most extensively with the J-chain. This may influence the multi-valent binding to surface-associated antigens and complement pathway activation. By comparison, the structure of the Fc fragment in the IgM monomer is similar to that of the pentamer, but is more dynamic in the Cμ4 domain.
Organizational Affiliation:
Structural Biology Science Technology Platform, The Francis Crick Institute, 1 Midland Road, London, NW1 1AT, UK.