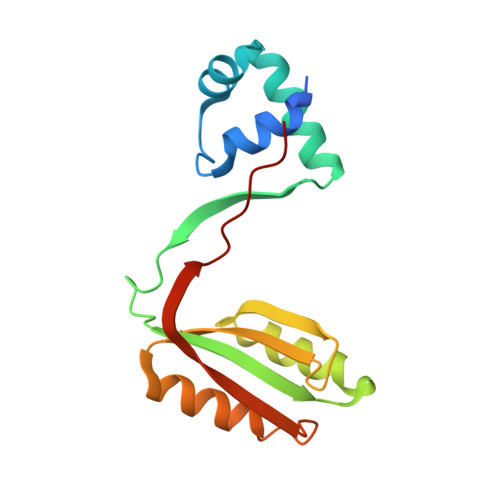
Global Lrp regulator protein from Haloferax mediterranei: Transcriptional analysis and structural characterization.
Matarredona, L., Garcia-Bonete, M.J., Guio, J., Camacho, M., Fillat, M.F., Esclapez, J., Bonete, M.J.(2024) Int J Biol Macromol 260: 129541-129541
- PubMed: 38244746
- DOI: https://doi.org/10.1016/j.ijbiomac.2024.129541
- Primary Citation of Related Structures:
8Q90 - PubMed Abstract:
Haloferax mediterranei, an extreme halophilic archaeon thriving in hypersaline environments, has acquired significant attention in biotechnological and biochemical research due to its remarkable ability to flourish in extreme salinity conditions. Transcription factors, essential in regulating diverse cellular processes, have become focal points in understanding its adaptability. This study delves into the role of the Lrp transcription factor, exploring its modulation of glnA, nasABC, and lrp gene promoters in vivo through β-galactosidase assays. Remarkably, our findings propose Lrp as the pioneering transcriptional regulator of nitrogen metabolism identified in a haloarchaeon. This study suggests its potential role in activating or repressing assimilatory pathway enzymes (GlnA and NasA). The interaction between Lrp and these promoters is analyzed using Electrophoretic Mobility Shift Assay and Differential Scanning Fluorimetry, highlighting l-glutamine's indispensable role in stabilizing the Lrp-DNA complex. Our research uncovers that halophilic Lrp forms octameric structures in the presence of l-glutamine. The study reveals the three-dimensional structure of the Lrp as a homodimer using X-ray crystallography, confirming this state in solution by Small-Angle X-ray Scattering. These findings illuminate the complex molecular mechanisms driving Hfx. mediterranei's nitrogen metabolism, offering valuable insights about its gene expression regulation and enriching our comprehension of extremophile biology.
Organizational Affiliation:
Departamento de Bioquímica y Biología Molecular y Edafología y Química Agrícola, Grupo Biotecnología de Extremófilos, Universidad de Alicante, San Vicente del Raspeig, Alicante, Spain.