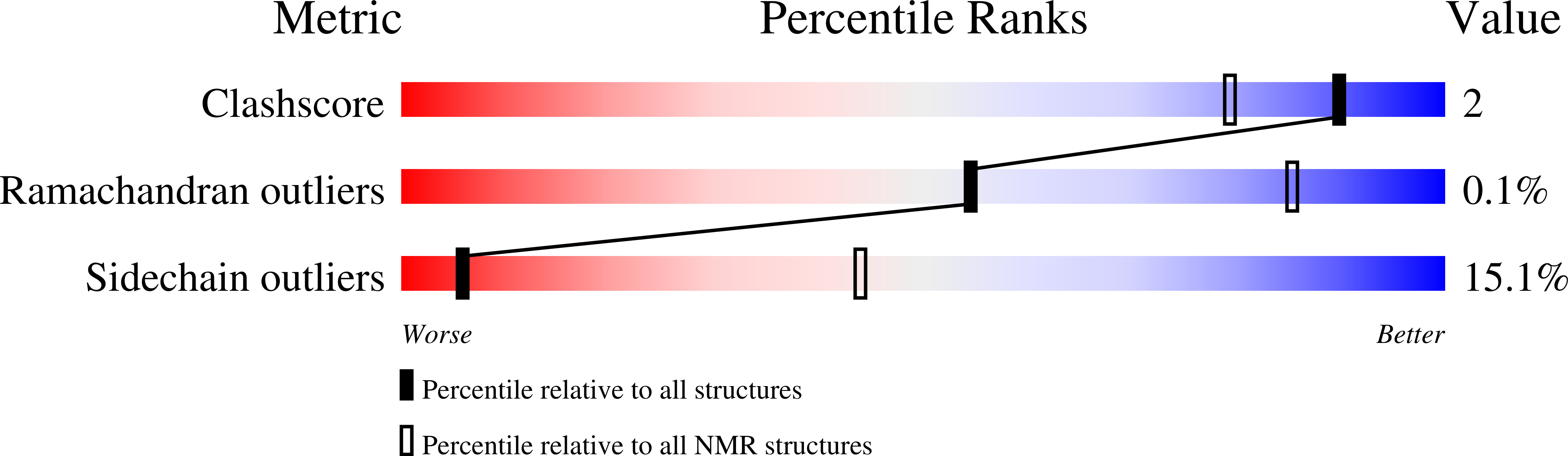RNA binding induces an allosteric switch in Cyp33 to repress MLL1-mediated transcription.
Blatter, M., Meylan, C., Clery, A., Giambruno, R., Nikolaev, Y., Heidecker, M., Solanki, J.A., Diaz, M.O., Gabellini, D., Allain, F.H.(2023) Sci Adv 9: eadf5330-eadf5330
- PubMed: 37075125
- DOI: https://doi.org/10.1126/sciadv.adf5330
- Primary Citation of Related Structures:
7ZEV, 7ZEW, 7ZEX, 7ZEY, 7ZEZ - PubMed Abstract:
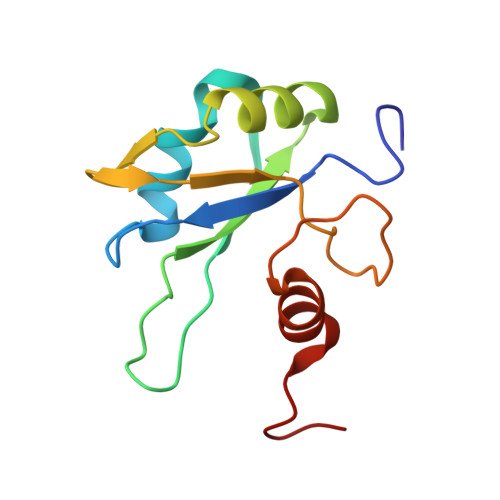
Mixed-lineage leukemia 1 (MLL1) is a transcription activator of the HOX family, which binds to specific epigenetic marks on histone H3 through its third plant homeodomain (PHD3) domain. Through an unknown mechanism, MLL1 activity is repressed by cyclophilin 33 (Cyp33), which binds to MLL1 PHD3. We determined solution structures of Cyp33 RNA recognition motif (RRM) free, bound to RNA, to MLL1 PHD3, and to both MLL1 and the histone H3 lysine N6-trimethylated. We found that a conserved α helix, amino-terminal to the RRM domain, adopts three different positions facilitating a cascade of binding events. These conformational changes are triggered by Cyp33 RNA binding and ultimately lead to MLL1 release from the histone mark. Together, our mechanistic findings rationalize how Cyp33 binding to MLL1 can switch chromatin to a transcriptional repressive state triggered by RNA binding as a negative feedback loop.
Organizational Affiliation:
Department of Biology, Institute of Biochemistry, ETH Zurich, 8093 Zurich, Switzerland.