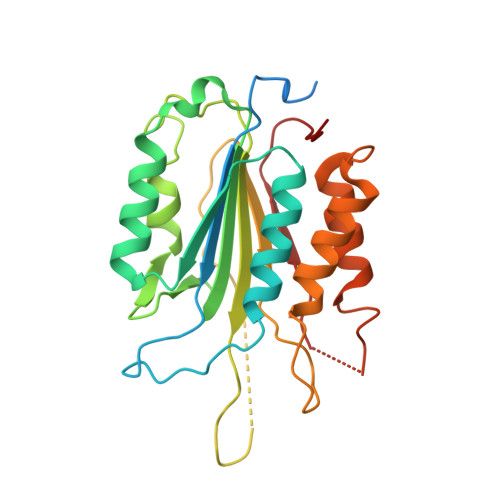
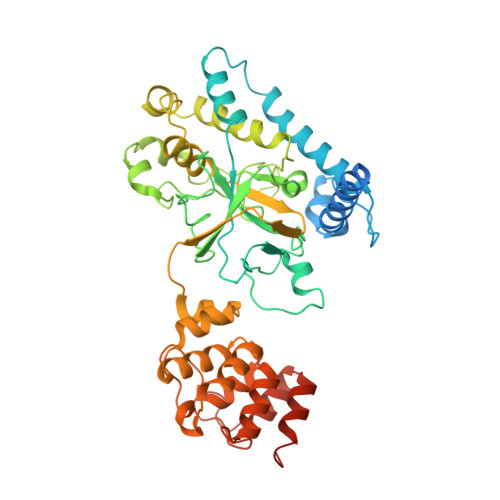
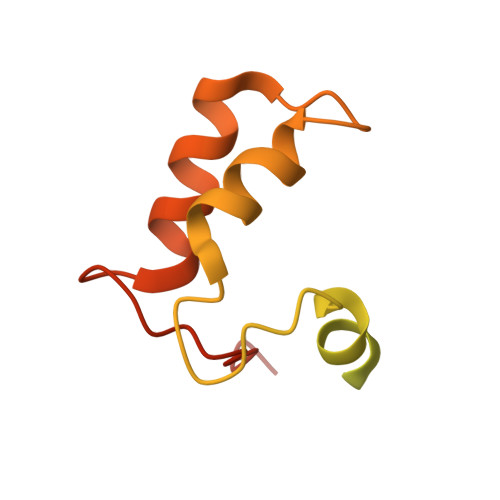
Structural insights into caspase ADPR deacylization catalyzed by a bacterial effector and host calmodulin.
Zhang, K., Peng, T., Tao, X., Tian, M., Li, Y., Wang, Z., Ma, S., Hu, S., Pan, X., Xue, J., Luo, J., Wu, Q., Fu, Y., Li, S.(2022) Mol Cell 82: 4712-4726.e7
- PubMed: 36423631
- DOI: https://doi.org/10.1016/j.molcel.2022.10.032
- Primary Citation of Related Structures:
7XN4, 7XN5, 7XN6 - PubMed Abstract:
Programmed cell death and caspase proteins play a pivotal role in host innate immune response combating pathogen infections. Blocking cell death is employed by many bacterial pathogens as a universal virulence strategy. CopC family type III effectors, including CopC from an environmental pathogen Chromobacterium violaceum, utilize calmodulin (CaM) as a co-factor to inactivate caspases by arginine ADPR deacylization. However, the molecular basis of the catalytic and substrate/co-factor binding mechanism is unknown. Here, we determine successive cryo-EM structures of CaM-CopC-caspase-3 ternary complex in pre-reaction, transition, and post-reaction states, which elucidate a multistep enzymatic mechanism of CopC-catalyzed ADPR deacylization. Moreover, we capture a snapshot of the detachment of modified caspase-3 from CopC. These structural insights are validated by mutagenesis analyses of CopC-mediated ADPR deacylization in vitro and animal infection in vivo. Our study offers a structural framework for understanding the molecular basis of arginine ADPR deacylization catalyzed by the CopC family.
Organizational Affiliation:
School of Medicine, Southern University of Science and Technology, Shenzhen 518055, Guangdong, China; Institute of Infection and Immunity, Taihe Hospital, Hubei University of Medicine, Shiyan 442000, Hubei, China.