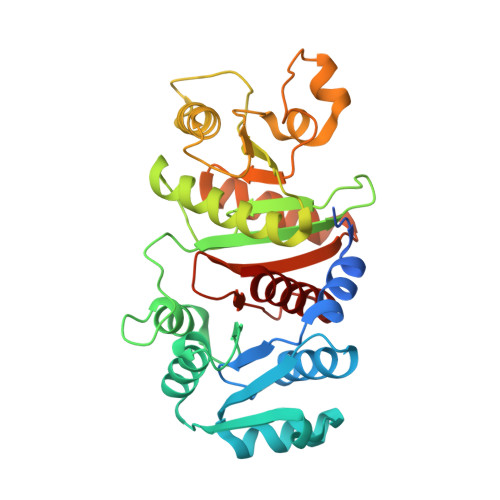
Crystal structure and molecular mechanism of phosphotransbutyrylase from Clostridium acetobutylicum .
Kim, S., Kim, K.J.(2021) J Microbiol Biotechnol 31: 1-9
- PubMed: 34584034
- DOI: https://doi.org/10.4014/jmb.2109.09036
- Primary Citation of Related Structures:
7VG9 - PubMed Abstract:
Acetone-butanol-ethanol (ABE) fermentation by the anaerobic bacterium Clostridium acetobutylicum has been considered a promising process of industrial biofuel production. Phosphotransbutyrylase (phosphate butyryltransferase, PTB) plays a crucial role in butyrate metabolism by catalyzing the reversible conversion of butyryl-CoA into butyryl phosphate. Here, we report the crystal structure of PTB from the Clostridial host for ABE fermentation, C. acetobutylicum , ( Ca PTB) at a 2.9 Å resolution. The overall structure of the Ca PTB monomer is quite similar to those of other acyltransferases, with some regional structural differences. The monomeric structure of Ca PTB consists of two distinct domains, the N- and C-terminal domains. The active site cleft was formed at the interface between the two domains. Interestingly, the crystal structure of Ca PTB contained eight molecules per asymmetric unit, forming an octamer, and the size-exclusion chromatography experiment also suggested that the enzyme exists as an octamer in solution. The structural analysis of Ca PTB identifies the substrate binding mode of the enzyme and comparisons with other acyltransferase structures lead us to speculate that the enzyme undergoes a conformational change upon binding of its substrate.
Organizational Affiliation:
School of Life Sciences, BK21 FOUR KNU Creative BioSesearch Group, Kyungpook National University, Daegu 41566, Republic of Korea.