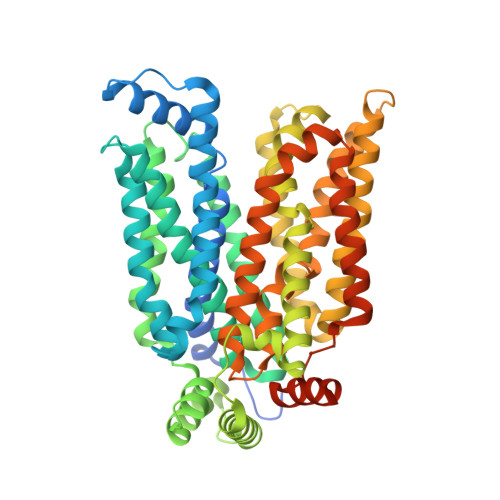
Molecular basis for inhibiting human glucose transporters by exofacial inhibitors.
Wang, N., Zhang, S., Yuan, Y., Xu, H., Defossa, E., Matter, H., Besenius, M., Derdau, V., Dreyer, M., Halland, N., He, K.H., Petry, S., Podeschwa, M., Tennagels, N., Jiang, X., Yan, N.(2022) Nat Commun 13: 2632-2632
- PubMed: 35552392
- DOI: https://doi.org/10.1038/s41467-022-30326-3
- Primary Citation of Related Structures:
7SPS, 7SPT - PubMed Abstract:
Human glucose transporters (GLUTs) are responsible for cellular uptake of hexoses. Elevated expression of GLUTs, particularly GLUT1 and GLUT3, is required to fuel the hyperproliferation of cancer cells, making GLUT inhibitors potential anticancer therapeutics. Meanwhile, GLUT inhibitor-conjugated insulin is being explored to mitigate the hypoglycemia side effect of insulin therapy in type 1 diabetes. Reasoning that exofacial inhibitors of GLUT1/3 may be favored for therapeutic applications, we report here the engineering of a GLUT3 variant, designated GLUT3exo, that can be probed for screening and validating exofacial inhibitors. We identify an exofacial GLUT3 inhibitor SA47 and elucidate its mode of action by a 2.3 Å resolution crystal structure of SA47-bound GLUT3. Our studies serve as a framework for the discovery of GLUTs exofacial inhibitors for therapeutic development.
Organizational Affiliation:
State Key Laboratory of Membrane Biology, Beijing Advanced Innovation Center for Structural Biology, Tsinghua-Peking Joint Center for Life Sciences, School of Life Sciences, Tsinghua University, Beijing, 100084, China.