Residue-Level Characterization of Antibody Binding Epitopes Using Carbene Chemical Footprinting.
Hogan, J.M., Lee, P.S., Wong, S.C., West, S.M., Morishige, W.H., Bee, C., Tapia, G.C., Rajpal, A., Strop, P., Dollinger, G.(2023) Anal Chem 95: 3922-3931
- PubMed: 36791402
- DOI: https://doi.org/10.1021/acs.analchem.2c03091
- Primary Citation of Related Structures:
7S7I - PubMed Abstract:
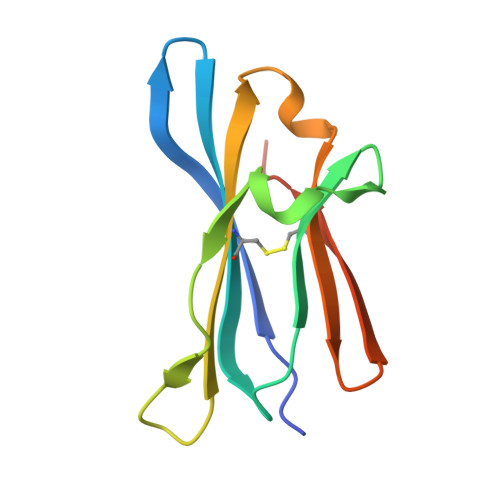
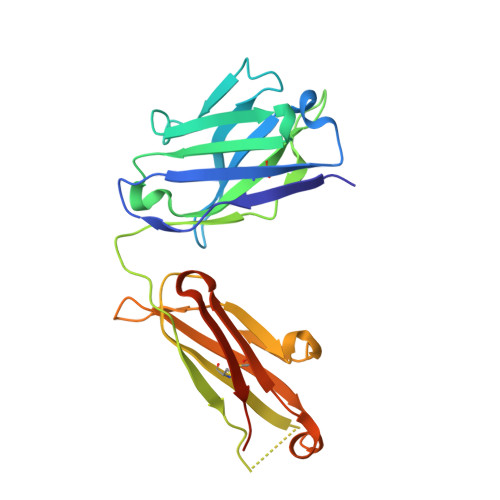
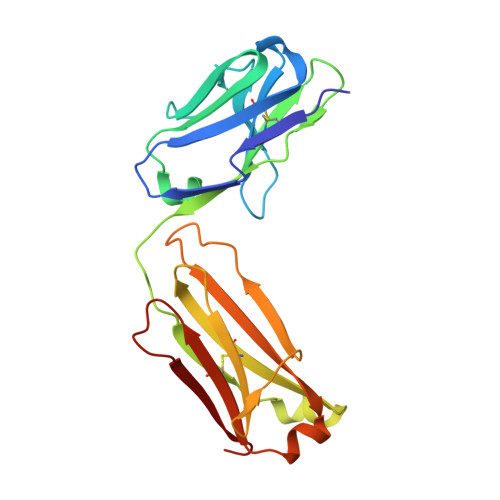
Characterization of antibody binding epitopes is an important factor in therapeutic drug discovery, as the binding site determines and drives antibody pharmacology and pharmacokinetics. Here, we present a novel application of carbene chemical footprinting with mass spectrometry for identification of antibody binding epitopes at the single-residue level. Two different photoactivated diazirine reagents provide complementary labeling information allowing structural refinement of the antibody binding interface. We applied this technique to map the epitopes of multiple MICA and CTLA-4 antibodies and validated the findings with X-ray crystallography and yeast surface display epitope mapping. The characterized epitopes were used to understand biolayer interferometry-derived competitive binding results at the structural level. We show that carbene footprinting provides fast and high-resolution epitope information critical in the antibody selection process and enables mechanistic understanding of function to accelerate the drug discovery process.
Organizational Affiliation:
Discovery Biotherapeutics, Bristol Myers Squibb, 700 Bay Road, Redwood City, California 94063, United States.